Page 160
Notes:
conferenceseries
.com
Volume 10, Issue 8 (Suppl)
J Proteomics Bioinform, an open access journal
ISSN: 0974-276X
Structural Biology 2017
September 18-20, 2017
9
th
International Conference on
Structural Biology
September 18-20, 2017 Zurich, Switzerland
Debarati Das Gupta et al., J Proteomics Bioinform 2017, 10:8(Suppl)
DOI: 10.4172/0974-276X-C1-0101
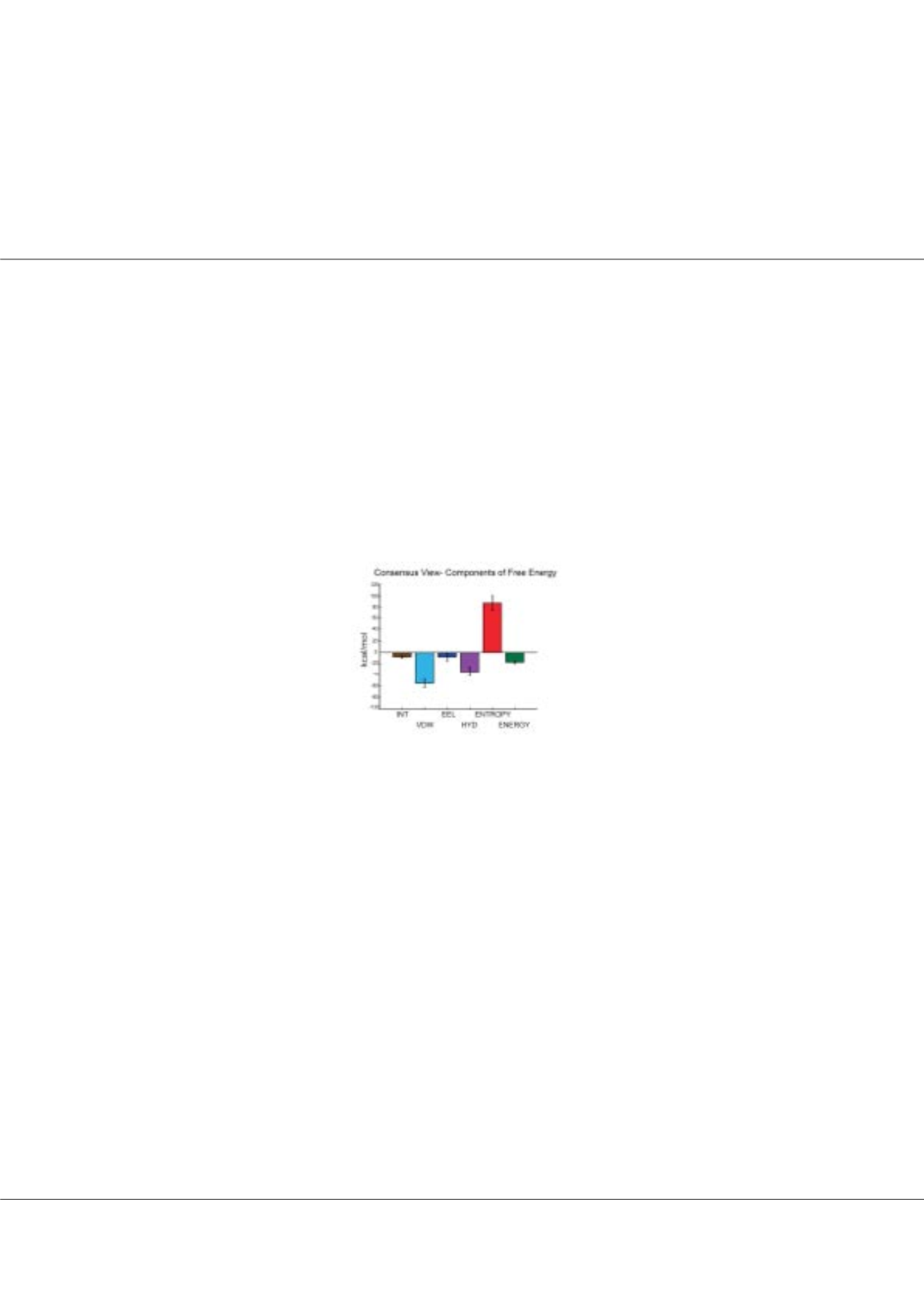
Consensus view of the energetics of protein folding studied on 35 proteins
Debarati Das Gupta, Mandalaparthy Varun
and
B Jayaram
Indian Institute of Technology Delhi, India
W
hat factors favor protein folding?This is a textbook question. Parsing the experimental free energies of folding/unfolding
into diverse enthalpic and entropic components of solute and solvent favoring or disfavoring folding is not an easy
task. In this study, we present a computational protocol for estimating the free energy contributors to protein folding semi-
quantitatively using ensembles of unfolded and native states generated
via
molecular dynamics simulations. We tested the
methodology on 35 proteins with diverse structural motifs and sizes and found that the calculated free energies correlate well
with experiment (correlation coefficient ~ 0.85), enabling us to develop a consensus view of the energetics of folding. As a more
sensitive test of the methodology, we also investigated the free energies of folding of an additional 33 single point mutants
and obtained a correlation coefficient of 0.8. A notable observation is that the folding free energy components appear to carry
signatures of the fold (SCOP classification) of the protein.
Biography
Debarati Das Gupta has obtained a Bachelor’s degree in Chemistry Honors which was conferred at St. Xavier’s College, Kolkata. She has done her Master’s in
Chemistry from Indian Institute of Technology Delhi. She has then joined the department as a PhD student under Prof. B Jayaram in July 2013. She is pursuing
PhD in the field of protein tertiary structure prediction, analysis of protein folding pathways and also energetic based studies using molecular dynamics simulations.
debarati@scfbio-iitd.res.in















