
Volume 10, Issue 8 (Suppl)
J Proteomics Bioinform, an open access journal
ISSN: 0974-276X
Structural Biology 2017
September 18- 20, 2017
Page 33
conference
series
.com
9
th
International Conference on
Structural Biology
September 18-20, 2017 Zurich, Switzerland
Yuri L Lyubchenko, J Proteomics Bioinform 2017, 10:8(Suppl)
DOI: 10.4172/0974-276X-C1-0100
Nanoscale structure and dynamics of centromere nucleosomes
Statement of the Problem:
Chromatin integrity is crucial for normal cell development.The cell division process is accompanied
by the segregation of replicated chromosome, and chromatin centromeres, specialized segments of chromosomes provide the
accuracy of the chromosomal segregation. If the centromere becomes damaged or removed, chromosomes segregate randomly
disrupting the cell division process. The centromeres are specifically recognized by kinetochores suggesting that centromeres
contain specific structural characteristics. However, these structural details and the mechanism underlying their highly specific
recognition remain uncertain.
Methodology &Theoretical Orientation:
In this study, we performed direct imaging of CENP-A nucleosome core particles by
time-lapse high-speed atomic force microscopy (AFM), enabling us to directly visualize the dynamics of CENP-A nucleosomes.
Nucleosomes used for evaluation of DNA wrapping around the histone core were assembled on a DNA substrate containing a
centrally positioned 601 motif.
Findings:
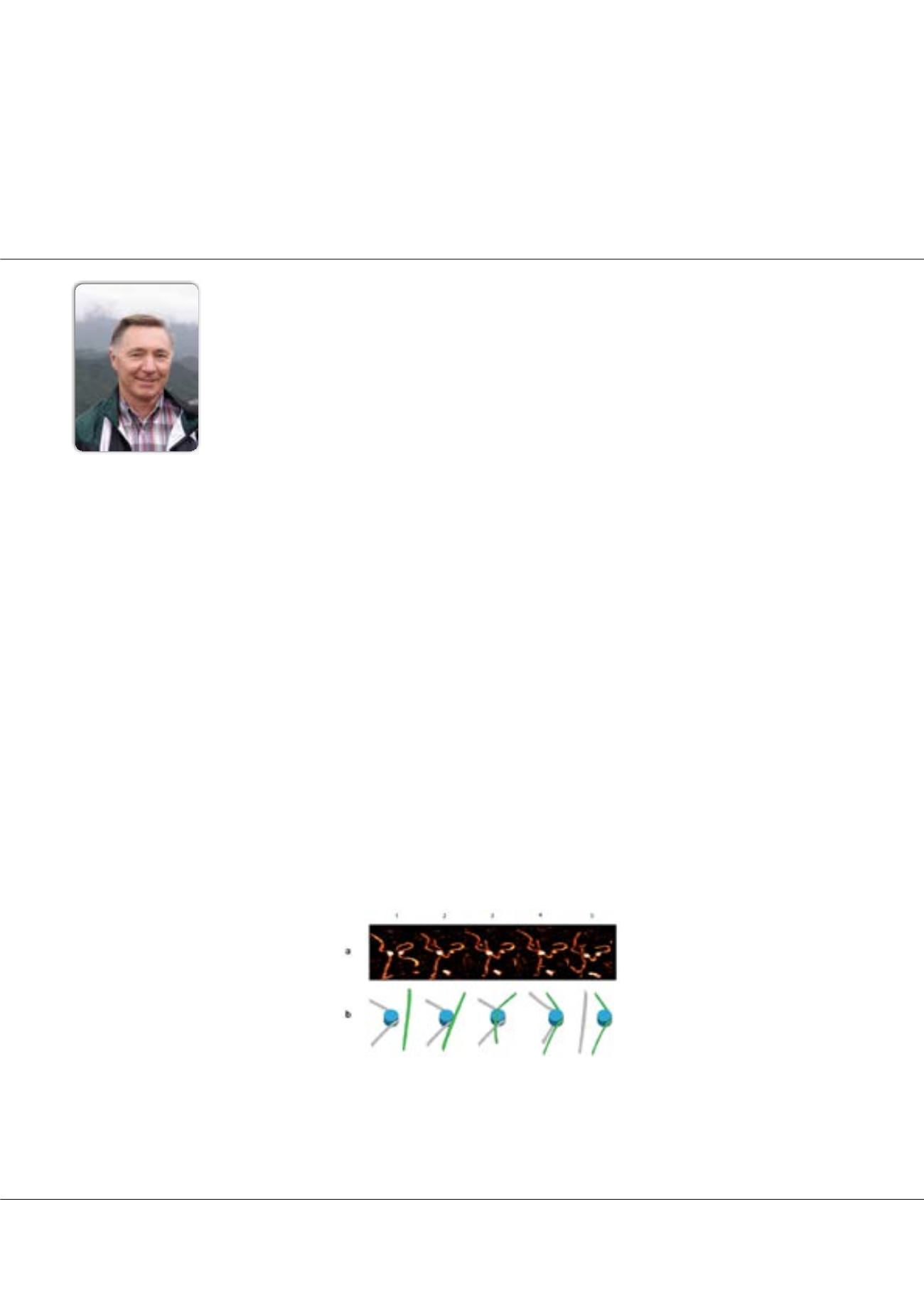
A broadly dynamic behavior of the DNA flanks was first revealed by analysis of AFM images acquired in ambient
conditions. Time-lapse imaging further identified the distinctive pathways unique to CENP-A-nucleosome dynamics that
are not shared by H3. The spontaneous unwrapping of DNA flanks can be accompanied by the reversible and dynamic
formation of loops with sizes equivalent to a single wrap of DNA. Translocation of CENP-A nucleosomes was observed, with
the formation of internal DNA loops along the nucleosome. This process was reversible, settling the core back to its starting
position. Additionally, the transfer of the histone core from one DNA substrate to another was visualized, as well as distinctive
splitting into sub-nucleosomal particles that was also reversible.
Conclusion & Significance:
Altogether, our data suggest that unlike H3, CENP-A is very dynamic, permitting its nucleosome
to distort freely and reversibly, which in turn allows a longer-term stability, which may play a critical role in centromere
integrity during mitosis and replication.
Biography
Yuri L Lyubchenko is the Professor of Pharmaceutical Sciences at the University of Nebraska Medical Center, Omaha, NE, USA. His research focuses on understanding
fundamental mechanisms underlying health and disease, which are key to developing new and more effective diagnostics and medications. This primarily basic research
allows him not only to identify new drug targets for small molecule drugs, it also leads to development of the nanotools and methods to discover novel approaches for
diagnostic, treatment and disease prevention and to more rapidly determine their efficacy at the molecular level.
ylyubchenko@unmc.eduYuri L Lyubchenko
University of Nebraska Medical Center, USA


















