Editorial Open Access
Metagenomics: A New Approach for Microbial Identification
Jian Yang*MOH Key Laboratory of Systems Biology of Pathogens, Institute of Pathogen Biology, Chinese Academy of Medical Sciences & Peking Union Medical College, China
- *Corresponding Author:
- Jian Yang
MOH Key Laboratory of Systems Biology of Pathogens
Institute of Pathogen Biology
Chinese Academy of Medical Sciences & Peking Union Medical College, China
Tel: 8610-6787-7735
E-mail:yangj@ipbcams.ac.cn
Received Date: June 19, 2012; Accepted Date: June 19, 2012; Published Date: June 20, 2012
Citation: Yang J (2012) Metagenomics: A New Approach for Microbial Identification. Air Water Borne Dis 1:e115. doi:10.4172/2167-7719.1000e115
Copyright: © 2012 Yang J. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Visit for more related articles at Air & Water Borne Diseases
Traditional microbiology focused on clonal cultures of single microorganism. However, it is now well established that majority of the microbes existing in the world could not be laboratory cultivated while playing essential roles in human life. Therefore, the newly emerged metagenomic method poses an important approach to explore the fantastic world of unculturable microorganisms. With the power of modern genomic techniques the metagenomic approach directly analyze the collective nucleotide contents of the entire microbial community (i.e. the metagenome), bypassing the need for isolation and lab cultivation of individual species [1].
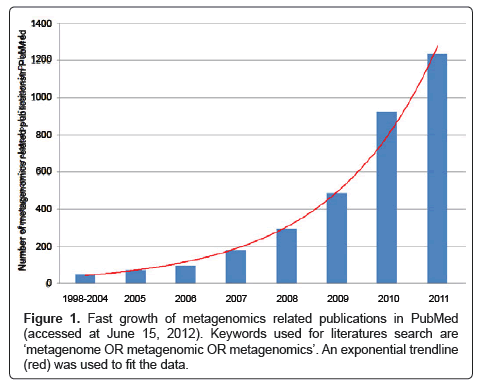
Though firstly proposed as early as 1998 [2], the metagenomic method was easily applicable only after the advent of next-generation sequencing (NGS) technologies, which have dramatically improved the speed and cost-effectiveness of DNA sequencing since 2005. In addition, the fast development of NGS platforms fueled the wide application of metagenomics in biomedical studies in recent years as reflected from the exponential increase in the number of related publications available in PubMed (Figure 1).
Though most of the former metagenomic studies focused on characterization of bacteria communities using 16S rDNA, many recent studies employed shotgun metagenomic strategy for screening all microorganism especially potential viral agents of human diseases. Using metagenomic approach on human tissue samples, Feng and coworkers discovered a previously unknown polyomavirus associated with Merkel cell carcinoma [3], and Palacios et al. also identified a new arenavirus related to a cluster of fatal transplant-associated diseases [4]. Metagenomic approach was further demonstrated to be a powerful complement to current methods for clinical diagnosis in accurate and parallel identification of various known viruses as well as novel variants from fecal [5-7] and nasal [8,9] specimen. For more information on applications of metagenomics in virus discovery refer to the recent review by Mokili et al. [10].
Nevertheless, some associated techniques need substantial improvements to facilitate further applications of metagenomic approach for microbial identification. Efficient methods for microbe enrichment are essential to circumvent the high human contamination as currently encountered in many studies. In addition, optimized bioinformatics tools for accurate binning, assembly and annotation from the metagenomic data are also urgently required.
References
- Chen K, Pachter L (2005) Bioinformatics for whole-genome shotgun sequencing of microbial communities. PLoS Comput Biol 1: 106-112.
- Handelsman J, Rondon MR, Brady SF, Clardy J, Goodman RM (1998) Molecular biological access to the chemistry of unknown soil microbes: a new frontier for natural products. Chem Biol 5: R245-R249.
- Feng H, Shuda M, Chang Y, Moore PS (2008) Clonal integration of a polyomavirus in human Merkel cell carcinoma. Science 319: 1096-1100.
- Palacios G, Druce J, Du L, Tran T, Birch C, et al. (2008) A new arenavirus in a cluster of fatal transplant-associated diseases. N Engl J Med 358: 991-998.
- Victoria JG, Kapoor A, Li L, Blinkova O, Slikas B, et al. (2009) Metagenomic analyses of viruses in stool samples from children with acute flaccid paralysis. J Virol 83: 4642-4651.
- Greninger AL, Runckel C, Chiu CY, Haggerty T, Parsonnet J, et al. (2009) The complete genome of klassevirus - a novel picornavirus in pediatric stool. Virol J 6: 82.
- Phan TG, Li L, O'Ryan MG, Cortes H, Mamani N, et al. (2012) A third gyrovirus species in human faeces. J Gen Virol 93: 1356-1361.
- Yang J, Yang F, Ren L, Xiong Z, Wu Z, et al. (2011) Unbiased parallel detection of viral pathogens in clinical samples by use of a metagenomic approach. J Clin Microbiol 49: 3463-3469.
- Lysholm F, Wetterbom A, Lindau C, Darban H, Bjerkner A, et al. (2012) Characterization of the viral microbiome in patients with severe lower respiratory tract infections, using metagenomic sequencing. PLoS One 7: e30875.
- Mokili JL, Rohwer F, Dutilh BE (2012) Metagenomics and future perspectives in virus discovery. Curr Opin Virol 2: 63-77.
Relevant Topics
Recommended Journals
Article Tools
Article Usage
- Total views: 8505
- [From(publication date):
December-2012 - Apr 04, 2025] - Breakdown by view type
- HTML page views : 3719
- PDF downloads : 4786