Decrypting the Code of Life DNA Sequencing
Received: 14-May-2018 / Accepted Date: 05-Jul-2018 / Published Date: 12-Jul-2018
Keywords: DNA sequencing; Genome project; Sanger sequencing
Introduction
Frederick Sanger pioneered DNA sequencing with his chain termination method in 1977 [1]. He and his team sequenced DNA samples by terminating the synthesis of DNA strand at each nucleotide using dideoxy dNTPs. This lead to the generation of many strands, each of different lengths corresponding to the different nucleotides on that strand. These were then separated and quantified using electrophoresis. Each band thus obtained on the gel was congruous to a nucleotide on the sample. The simplicity, availability and the accuracy of the method helped Sanger win his second Nobel Prize. There were researchers like Maxam et al. [2] that came up with other innovative methods like chemical modification with base-specific cleavage but the efficiency of Sanger sequencing helped it mark its place in history. Slabs of gel were replaced with automated multicapillary electrophoresis which combined with better fluorescent dye detection sequenced hundreds of strands simultaneously and making Sanger sequencing, the industry standard.
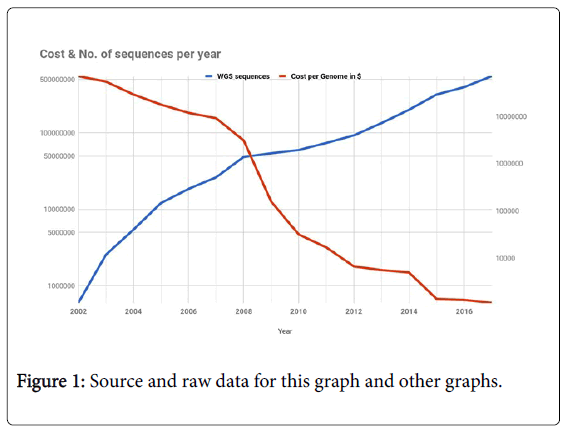
From the last quarter of the 20th century to the early 2000’s, Sanger sequencing was the go-to aid for almost all researchers. When the idea of Human Genome Project, the ambitious plan to sequence all 3 billion base pairs of a human genome was drafted in 1990’s, automated Sanger’s method was the primary choice to shine the flashlight unto the secret of life which was carefully vaulted in the nucleus. The Human Genome Project (HGP) was started in 1990, it was estimated to end in 15 years, but the first working draft was produced in 2000 and by 2003 [3], 2 years ahead of schedule, HGP was complete [4]. A private company Celera Genomics [5] also produced a similar sequence at the same time. HGP cost $3 billion dollars, 13 years and thousands of scientist to sequence a human genome, which as of 2018 would take hardly cost $ 1000 and few hours. In the last decade after the HGP, a new class of gene sequencing technologies, which referred as the second generation, or the Next Generation Sequencing (NGS) technology have not just decreased the cost, but also increased the accessibility. The following graph encapsulates NGS and its growth (Figure 1).
NGS: Origin, evolution and types
If automated Sanger sequencing was the first generation, the NGS or High Throughput Sequencing is the second gen. NGS is an umbrella term of different technologies that rely on varying chemistries to decrypt the code of life. The boom in these NGS technologies came in the middle of the first decade of the 21st century. 454 sequencing platform by Roche became commercially available by 2005 [6]. While these NGS were better than Sanger in terms of accuracy and cost, they struggled with reading lengths. 454 were closely followed by ABI SOLiD sequencing, Life ion torrent, Illumina’s genome analyzer and others. Increased hard drive capacity, processing power, network bandwidth and the general boom of computer technology lead to a rapid shift from studies based on few-loci to the genome-wide analysis studies. NGS sequenced DNA by first breaking it into short samples of readable length, annealing them, adding an oligonucleotide primer, amplifying them, sequencing them and eventually stitching them together by matching the overlaps in the short sequences [7]. All the short sequences are read simultaneously and hence is also called as Massively parallel sequencing. The primary mode of working for NGS involves 3 broad steps.
1. Template creation and amplification.
2. Sequencing and imaging.
3. Alignment and assembly of genomic data.
Let’s look further into the different types of NGS and how these steps fall into their protocol.
Roche 454/Pyrosequencing: Instead of dideoxy dNTPs, this method relies on detection of pyrophosphate that is released on the incorporation of NTPs [8]. Emulsion PCR is used to capture denatured DNA strand on amplification beads. An enzyme called ATP luciferase generates light when pyrophosphate (PPi) is released on induction of dNTP in the denatured DNA strand. This light is different for every dNTP (dATP, dTTP, dCTP, dGTP) and is recorded with each new round of dNTP wash.
Illumina: It was developed by Solexa which was acquired by Illumina Inc in 2007. The prepared sample is used to make small clusters on the flow cell using bridge amplification. The annealed strand is synthesized in double strand using fluorescent dNTPs on flowcells. Each dNTP gives a different color when analyzed by a laser. This color is recorded by a camera and analyzed by computers. Illumina and 454 are together also called as sequencing by synthesis method.
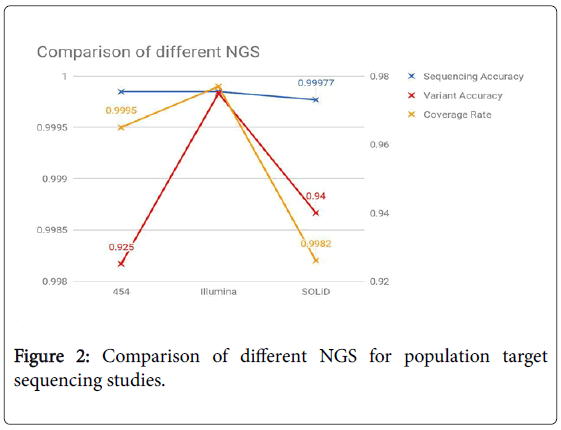
SOLiD: This method uses ligation and two base sequencing. The first step is similar, DNA is denatured, oligos and adapters are added and the strands are amplified using hybridized beads. An 8 base probe (first 2 bases are ligation site, next 3 are cleavage site and the last 3 are the fluorescent site) is used to find the nucleotide on the strand. 5 cycles of ligation, detection, and cleavage is performed for each sequence tag. SOLiD stands for Sequencing by Oligonucleotide Ligation and Detection (Figure 2).
NGS outperform Sanger on many fronts. Chemical reaction and signal detection are two individual steps in Sanger but are combined into one in most NGS. Sanger can process one read of a longer length but NGS, on the other hand, can parallelly sequence hundreds of reads. NGS is not just inexpensive, but also easy to use and faster. The domination of NGS has led to the better variant identification and more exhaustive understanding of epigenomes and transcriptomes. The decrease in the cost per base pair and the sequencing of a large number of diverse genomes has given us a more coherent understanding of genes and their impact on our lives.
Applications
The need for DNA sequencing is rising rapidly. It’s been used across a wide range of diverse verticals like evolution, epidemiology, forensic, diagnosis, genetic engineering, archaeology, precision medicine, population genetics etc. The ability to completely understand the genetic makeup of an organism gives researchers the ability to not only compare it with other organisms to identify phylogenetic juxtaposition but also find individual differences that lead to organism wide variation [9-11]. The biggest use of DNA sequence is perhaps to understand our genes and how they make us sick.
Another important rationale behind DNA sequencing is gene therapy and gene engineering. You can certainly not edit the code of life if you can’t read it. The improved accessibility and the ease of use have driven metagenomics, also known as community genomics which refers to the study of a genetic sample which is directly collected from the environment. The flow of genes and their expression rate is an useful information to deduct their common evolutionary tracts. Single-molecule sequencing is also used to detect complex structural variations [12]. Large-scale population study like the All of us project by NIH [13] and the 100,000 Genome Project by the NHS [14] have been possible because of inexpensive sequencing tools. These projects aim to get more insight on gene and disease association and use the data to make precision medicine, that is customized to the genetic makeup of the patients. The exact sequence of nucleotide in a gene can be used to predict the protein it would transcribe by using computer models. Along with genes, sequencing also helps researchers understand the role regulatory devices like promoters, enhancers, silencers in gene expression. The impact of DNA sequencing is spread across all domains of genetic and evolutionary science.
Oxford nanopore
Oxford Nanopore in a DNA sequencing platform that reads a single strand of DNA by measuring the change in electric current when it passes through biological pores embedded in a membrane. Strands are run through a pore using motor proteins like helicase. Conceptualized in 1980, it took until 2012 for it is materialized. As of 2018, products using Nanopore technology are available for research purposes, but yet to be released commercially.
Two things that separate Oxford Nanopore from traditional NGS is the long read lengths (in excess of 800,000 bps [15] and accessibility. While conventional sequencing machines are as big as a large oven, Nanopore is a USB operated, keychain-sized device. A 100g sequencing device can not only be carried on the ground for field work but also be taken to remote places where transportation is a major challenge. In 2016, researchers were able to sequence the Ebola virus at the heart of the epidemiology nightmare in West Africa [16]. Nanopore was also used to decipher the smallest centromere in humans [17], on the Y chromosome with the help of BAC (bacterial artificial chromosome), a feat no other NGS could do.
Oxford Nanopore still lags behind other platforms in terms of accuracy in base calling. The technology is still new and being developed for commercial purposes. Oxford Nanopore has several products like MinION and PromethION which have democratized DNA sequencing. The unmatched portability and inexpensive run promises to one day, bring sequencing to every lab on earth.
Future
There is a new wave of TGS or Third generation sequencing tools that have started to flood the market [18]. Nanopore and Pacific Biosciences single polymerase sequencing have been leading this wave. Advancement in cloud computing and better processors is further going to decrease the cost and improve read length and accuracy. Companies like 23 & me and Ancestry.com which commercially genotype their user’s DNA sample have seen a massive year on year growth. From Genotyping, the next very likely frontier for these companies will be to offer large-scale Whole Exome and Whole Genome Sequencing. More genomic data will lead to better prediction and of disease and its prognosis and eventually help providers chart a better course for recovery. Google recently released their new Deep Learning algorithm (Artificial Intelligence) called Deep Variant that helps in reconstructing full genomes from raw HTS data and apply Machine Learning to find SNPs.
From microbiologist, taxonomist, archeologist, evolutionary and marine biologist, everyone is using DNA sequencing to understand the flow of life at the most microscopic level. The idea of the most intimate secrets of an organism being sequenced with such ease raises a lot of privacy and surveillance concerns. Will we be moving into a world where anyone with a sequencer can steal and read anyone’s genome? While privacy concerns are a valid worry, the right amount of caution has to be exercised while moving forward. Looking at the exponential graph of growth, the future we await is closer than it seems. Third generation sequencing platforms and Oxford Nanopore combined with Artificial Intelligence has a lot of offer to mankind. Just like Sanger sequencing, this amalgamation of technology and biology is a promising insight into a future where decrypting the code of life will be a click away.
References
- Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. ​Proceedings of the National Academy of Sciences of the United States of America ​74: 5463-5467.
- Maxam AM, Gilbert W (1977) A new method for sequencing DNA. Proc Natl Acad Sci USA 74: 560-564.
- Lander ES, Linton LM, Birren B, Nusbaum C, Zody MC, et al. (2001) Initial sequencing and analysis of the human genome Nature, Nature 409: 860-921.
- Human Genome Sequencing Consortium I (2004) Finishing the euchromatic sequence of the human genome Nature 431: 931-945.
- Venter JC, Adams MD, Myers EW, Li PW, Mural RJ, et al. (2001) The Sequence of the Human Genome. Science 291: 1304-1351.
- Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, et al. (2005) Genome Sequencing in Open Microfabricated High Density Picolitre Reactors. ​Nature ​437: 376-380.
- Metzker ML (2009)Â Sequencing technologies-the next generation. Nature Reviews Genetics 11: 31-46.
- Rothberg JM, Leamon JH, (2008) The development and impact of 454 sequencing. Nature Biotechnology 26: 1117-1124. ​
- Harismendy O, Ng PC, Strausberg RL, Wang X, Stockwell TB (2009) Evaluation of next generation sequencing platforms for population targeted sequencing studies. Genome Biology 10: R32.
- Quail M, Smith ME, Coupland P, Otto TD, Harris SR, et al. (2012) A tale of three next generation sequencing platforms: comparison of Ion torrent, pacific biosciences and illumina MiSeq sequencers. BMC Genomics 13: 341.
- Metzker ML (2009) Sequencing technologies the next generation. Nature Reviews Genetics 11: 31-46.
- Sedlazeck FJ, Rescheneder P, Smolka M, Fang H, Nattestad M, et al. (2018) Accurate detection of complex structural variations using single-molecule sequencing. Nature Methods 15: 461-468.
- Jain M, Koren S, Miga KH, Quick J, Rand AC, et al. (2018) Nanopore sequencing and assembly of a human genome with ultra-long reads. Nature Biotechnology 36: 338-345.
- Quick J, Loman NJ, Duraffour S, Simpson JT, Severi E, et al. (2016) Real-time, portable genome sequencing for Ebola surveillance. Nature 530: 228-232.
- Jain M, Olsen HE, Turner DJ, Stoddart D, Bulazel KV (2018) Linear assembly of a human centromere on the Y chromosome. Nature Biotechnology 36: 321-323.
- Schadt EE, Turner S, Kasarskis A (2010) A window into third-generation sequencing. Human Molecular Genetics 19: 227-240.
- Korlach J, Bjornson KP, Chaudhuri BP, Cicero RL, Flusberg BA, et al. (2010) Real-Time DNA Sequencing from Single Polymerase Molecules. Methods in Enzymology 472: 431-455.
- Poplin R, Chang PC, Alexander D, Schwartz S, Colthurst T, et al. (2016) Creating a universal SNP and small indel variant caller with deep neural networks. Cold Spring Harbor Laboratory. bioRxiv 092890.
Citation: Borana RA (2018) Decrypting the Code of Life DNA Sequencing. J Genome 1: 103.
Copyright: © 2018 Borana RA. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Share This Article
Open Access Journals
Article Usage
- Total views: 3470
- [From(publication date): 0-2018 - Mar 31, 2025]
- Breakdown by view type
- HTML page views: 2650
- PDF downloads: 820