Research Article Open Access
Cloning and Analysis of N-Acetyltransferase 9 Genes in Yellow Catfish Pelteobagrus fulvidraco
| Ming-Ming Han1,2, Jian-Guo Lu1, Le-Wang1, Li-Na-Peng1, Shahidd Mahboob3,4, Khalid A Al-Ghanim4 and Xiao-Wen Sun1* | |
| 1Heilongjiang River Fisheries Research Institute, Chinese Academy of Fishery Sciences, Harbin, China | |
| 2Shanghai Ocean University, College of Fisheries and Life Science, Shanghai, China | |
| 3Department of Zoology, College of Science, King Saud University, Riyadh, Saudi Arabia | |
| 4Department of Zoology, GC University, Faisalabad, Pakistan | |
| *Corresponding Author : | Xiao-Wen Sun Heilongjiang River Fisheries Research Institute Chinese Academy of Fishery Sciences, Harbin, China Tel: +86-451-84869349 Fax: +86-451-84604803 E-mail: sunxw2002@163.com |
| Received: February 13, 2016 Accepted: March 03, 2016 Published: March 10, 2016 | |
| Citation: Han MM, Lu JG, Wang L, Peng LN, Mahboob S, et al. (2016) Cloning and Analysis of N-Acetyltransferase 9 Genes in Yellow Catfish Pelteobagrus fulvidraco. Biochem Physiol 5:202. doi: 10.4172/2168-9652.1000202 | |
| Copyright: © 2016 Han MM, et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Visit for more related articles at Biochemistry & Physiology: Open Access
Abstract
N-acetyltransferase 9 (NAT9) is an important reproduction-related gene in fish. In this study, we cloned the fulllength NAT9 cDNA sequence from yellow catfish Pelteobagrus fulvidraco. P. fulvidraco NAT9 cDNA is 1253 bp, which including non-coding region (584 bp) and open reading frame (669 bp). NAT9 gene encodes a 222 amino acidsprotein which shares high homology with NAT9 in four species: Ietalurus punetaus (91%), Astyanax mexicanus (83%), Danio rerio (81%) and Lepisosteus oculatus (81%). NAT9 protein is 25.7899 kD, 4.86 theoretical isoelectric point and C1124H1759N311O359S13 as revealed by computer-assisted analysis. Tissue transcription profile analysis indicated that the catfish NAT9 gene is generally but differentially expressed in the detected tissues, including gonads, brain, liver, kidney, spleen, intestine, blood, gills, muscle and heart. Identification and tissue distribution of yellow catfish NAT9 genes provided initial step towards understanding their biological roles in yellow catfish.
| Keywords |
| N-acetyltransferase 9; Pelteobagrus fulvidraco; Gene clone; Bioinformatics analysis |
| Introduction |
| N-acetyltransferases (NATs) are well characterized xenobioticmetabolizing enzymes which catalyze the acetyl Coenzyme A (AcCoA)- dependent N-, O- and N,O-acetylation of aromatic amines and hydrazines. They played important endogenous roles, as well as having potential as novel targets for pharmacological intervention. NAT9 is a member of NATS which has N-acetyltransferase, acyltransferase and transferase activities. NAT9 has important biological function, due to the neutral protein electropositivity [1]. NAT acetylates wide range of drugs and xenobiotics into prominence, as acts in the polymorphic metabolism of the anti-tubercular agent isoniazid [2]. |
| Latest studies reported that NAT9 was highly expressed in human brain, gonad, and specifically expressed in the heart, spleen, gonad of adult mouse. Whole-mount in situ hybridization showed that NAT9 specifically expressed in E7.5 and E8.5 mouse embryo brains and in HH10 stage chicken embryo brain. These results suggest that NAT9 may play important roles in the development of embryonic brain and adult brain and gonad’s function [3]. As above reports, NAT9 is an important reproduction related gene. Although NAT9 gene was found has expression in human [4], rat [5], cattle [6] and other animals, this gene expression in yellow catfish has not been reported yet. Whether NAT9 gene is associated with the reproductive traits in yellow catfish is still unknown. |
| Yellow catfish Pelteobagrus fulvidraco, an omnivorous freshwater fish, is regarded as a good candidate for freshwater culture in China for its delicious meat and high market value [7]. In this study, we obtained the full-length cDNA sequence of NAT9 gene, and analyzed tissue expression and putative proteins through bioinformatics approaches. The results of our study provide the basic information of NAT9, and illuminate the molecular mechanism of this gene in P. fulvidraco. |
| Materials and Methods |
| RNA isolation |
| The P. fulvidraco were obtained from the Chinese Academy of Fishery Sciences P. fulvidraco Breeding Engineering Center. The yellow catfish were 2 years old, 60.6 ± 2.7 g weight and 18.5 ± 3.4 cm length. The extraction of total RNAs from the yellow catfish was conducted using the Trizol (Invitrogen, China) technique following the manufacturer’s instructions. The total RNA concentration was determined by measuring the absorbance at OD260. RNA integrity was checked by electrophoresis. The total RNA was reversetranscribed into cDNA using the aM-MLV RTase cDNA Synthesis Kit (TaKaRa, Japan). |
| NAT9 cloning |
| Every tissue total RNA used cDNA reverse treasure article number for RR047A transcription kit and used minus 20 degrees to put on cDNA template. According to Ietalurus punetaus in Genbank madtoms NAT9 (serial number DQ353801.1) cloning NAT9 cDNA partial sequence. After the sequence by Heilongjiang River Fisheries Research Institute of products research, development of genomics and molecular breeding research of yellow catfish are found in ninety-nine percent of similarity in the method of the transcriptome gene sequence and gene sequence designed primers. We designed the PCR primers NAT9-F1, NAT9-R1, NAT9-F2, NAT9-R2, NAT9-F3, NAT9-R3, NAT9-F4, NAT9-R4, NAT9-F5 and NAT9-R5 to clone the NAT9 cDNA partial sequence (Table 1). |
| PCR was conducted for 5 min at 94°C followed by 35 cycles at 94°C for 30 s, 56°C for 30 s, and 72°C for 30 s. The final extension step was performed at 72°C for 2 min. Each of the 20 μL reaction mixtures contained 1.5 mM of MgCl2, 0.2 mM of dNTP, 0.3 mM of each of the primers, 1 U of Taq DNA polymerase (TaKaRa, Japan), and 5 ng of cDNA. Amplicons of expected sizes were purified using an Agarose Gel DNA Purification Kit (TaKaRa, Japan), and then subcloned into the pMD-18T cloning vector (TaKaRa) connection at 16°C for the night. Tiangen Top 10 into competent E. coli in LB Amp tablet on 37°C, or 230 r/min, Pick the bacteria, expand training and gas-bacilli PCR detection, expand training and gas-bacilli PCR detection. PCR products were 0.5% gel electrophoresis detection of target DNA in bacteria. Positive clones containing inserts of an expected size were sequenced using M13 primers and sequenced at BGI (Shanghai, China). The sequencing results on NCBI homology retrieval. |
| Sequence analysis of NAT9 |
| Deduced amino acid sequences of NAT9 cDNA were analyzed using the software Vector NTI Advance 11. NAT9 sequences from different organisms were obtained using the NCBI BLAST search program. The cDNA sequence and deduced amino acid sequence from Yellow catfish were analyzed using the BLAST algorithm (http://www.ncbi. nlm.nih.gov/blast). Translation and protein analyses were performed using ExPASy tools (http://us.expasy.org/tools/). A multiple sequence alignment was created with ClustalW (http://www.ebi.ac.uk/clustalw/). Protein sequences retrieved from the public database were used for ORF and domain searches; alignment; and phylogenetic reconstruction. ORFs were predicted using Open Reading Frame Finder (http://www. ncbi.nlm.nih.gov/gorf/gorf.html), and signal peptides and fibrinogenlike domain (FBG) were identified by the NCBI conserved domain feature of blastp (http://www.ncbi.nlm.nih.gov/BLAST) and by the Simple Modular Architecture Research Tool (SMART) (http://smart. emblheidelberg.de). |
| Tissue expression of NAT9 gene |
| NAT9 gene was in accordance with full design relative fluorescence quantitative PCR primers for NAT9-F5 and NAT9-R5 (Table 1). Primers were 125 bp amplification length. Primers respectively had actins-F and actins-R. The length of amplified fragment is 200 pb. It can enter the yellow catfish beta actin genes for internal reference. RT-PCR reverse transcription using the reverse transcription kit ABI. The PCR reaction was performed in a 25 lL volume with a SYBR Premix Ex Taq™ Kit (TaKaRa, Japan). Experiment according to the instructions, the reaction fluorescence quantitative instrument (ABI 7500) on the conduct for analysis using 2 μM of each specific primer, and 1 lL of cDNA using the following procedure: initial denaturation at 95°C for 2 min followed by 40 cycles of amplification (95°C for 10 s and 55°C for 30 s). Experimental analysis need to â�?³â�?³CT. Relative quantitative calculation method: relative quantitative is equal to 2â�?³â�?³CT multiplied by 100%. |
| To determine NAT9 gene expression in, samples of 9 tissues including brain, muscle, kidney, gill, intestine, heart, brain, spleen and blood from various healthy male and female yellow catfishes were 2 years old, 60.6 ± 2.7 g weight, 18.5 ± 3.4 cm length, a total of 30 and some channel catfishes as control were isolated, pooled, and flashfrozen in liquid nitrogen. The yellow catfish Tissues were homogenized under liquid nitrogen using a mortar and pestle, and stored at −80°C until RNA extraction. |
| Phylogenetic analysis |
| The amino acid sequence and coding regions of NAT9 gene were deduced using the Primer 5 software, NAT9 gene were excerpts from GenBank, number for KT778803 GenBank landed. Sequences of NAT9: Ietalurus Punetaus, Astyanax mexicanus zebra, Danio rerio, Esox Lucius, Oncorhynchus mykiss, Lepisosteus oculatus, Reochromis spp., Clupea harengus, Astatotilapia burtoni, Oryzias latipes, Neolamprologus brichardi, Fundulus heteroclitus, Xiphophorus maculatus,Fugu rubripes and Poecilia Formosa retrieved from databases were aligned using the ClustalW2 program (http://www.ebi.ac.uk/Tools/clustalw2/). Phylogenetic trees were constructed using the neighbour-joining (NJ) method based on the deduced full-length amino acid sequences with 1,000 bootstrapping replications within the Molecular Evolutionary Genetics Analysis (MEGA 4.0) package. |
| Bioinformatics analysis |
| The physicochemical properties, protein domains, secondary structures, secretory proteins, subcellular localization signals, transmembrane regions, and 3D structure models of the putative proteins of NAT9 were predicted and analyzed using online bioinformatics tools, as listed in Table 2. To strengthen the conclusions, more than one bioinformatics tool was used in the prediction of secretory proteins, subcellular localization signals, and transmembrane regions for the protein. The sequence alignment of amino acids was performed by using the BLASTX program, which is available on the NCBI website. The amino acid sequences of HsHDR1 and HsHDR2 were aligned, and the phylogenetic tree was constructed with neighborjoining criteria by the MEGA 6.0 software. |
| Statistical analysis |
| All data obtained from the qRT-PCR were calculated to the value of 2-â�?³â�?³CT and differences were evaluated statistically using a t-test in SPSS 20.0 software (IBM, Chicago, USA). The relative expression levels of each gene in different tissues were calculated to the 2-â�?³â�?³CT of the mean and the standard error (SE) from the three replications. Statistical significance was set with a P value less than 0.01. |
| Results |
| Phylogenetic analysis of the NAT9 gene |
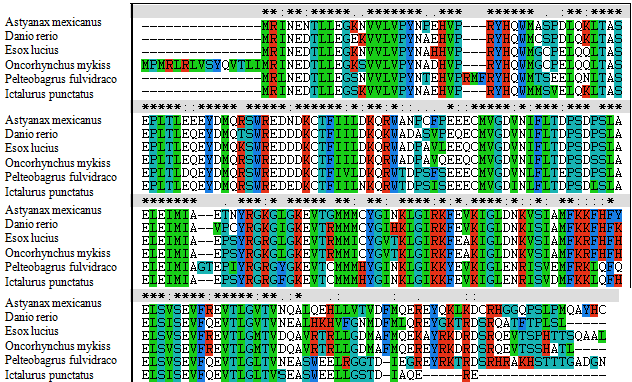
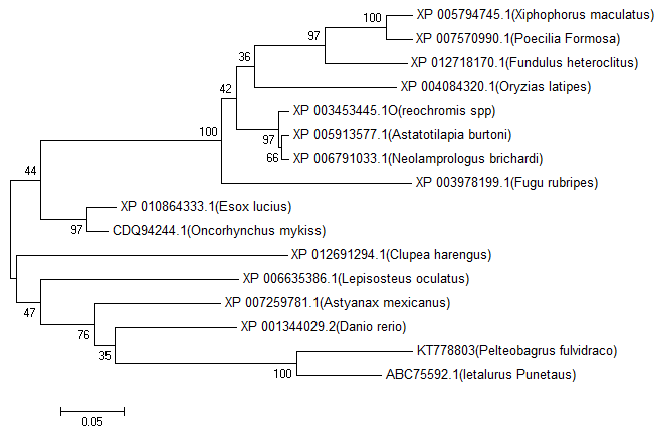
| A phylogenetic tree was constructed based on the amino acid sequences of selected NAT9 using the neighbour-joining (NJ) method (Figures 1 and 2). Data were analyzed by Poisson’s correction, and gaps were removed by complete deletion. Neolamprologus brichardi, Astatotilapia burtoni and Reochromis spp. belong to Perciformes; Fundulus heteroclitus, Oryzias latipes, Xiphophorus maculatus and Poecilia formosa belong to Cyprinodontiformes; Fugu rubripes of Tetraodontiformes and yellow catfish NAT9 amino acids have different branches. The results showed that the molecular NAT9 evolutionary relationships and the corresponding species are known similar evolutionary relationships. The NAT9 gene encodes a protein of 222 amino acids which shares high homology with the NAT9 from four species: Ietalurus punetaus (91%), Astyanax mexicanus (83%), Danio rerio (81%) and Lepisosteus oculatus (81%) (Figure 2). |
| Tissue expression of NAT9 gene |
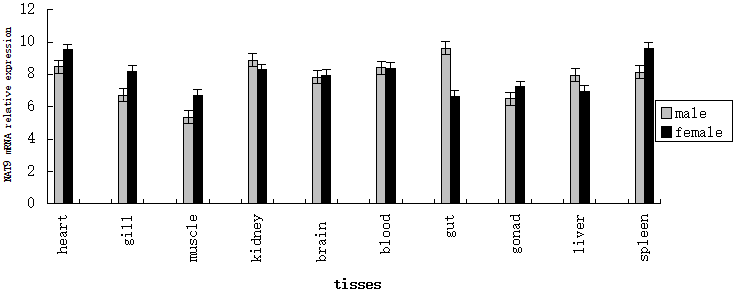
| Quantitative real-time PCR was used to determine tissue distribution of NAT9 gene expression in both male and female P. fulvidraco. NAT9 genes were widely expressed in all 10 tissues (gonads, brain, liver, kidney, spleen, intestine, blood, gills, muscle and heart). NAT9 gene expression in the male gut and liver were higher than in female. NAT9 gene expression in the female heart, gill, muscle and spleen were higher than males (Figure 3). |
| Analysis of the physicochemical properties |
| We analyzed the physicochemical properties of the putative NAT9 protein using the ProtParam (http://www.au.Expasy.Org/tools/ protparam.html). The molecular weight of NAT9 protein was 25.7899 kD and 4.86 theoretical isoelectric point. The negatively charged amino acid residues (Asp + Glu) was 40, and positively charged amino acid residues (Arg + Lys) was 26. The instability coefficient of NAT9 protein was 48.15, indicating that the protein was unstable. Theoretical derivation half-life of NAT9 protein was about 30 h. The fat coefficient was 0.669 and grand average of hydrophobic was 71.94. Amino acid content: Glu 12.2%, Leu 9.0%, Thr 7.7%, Arg 7.2%, Gly 6.8%, Ser 6.3%, Asp 5.9%. The hydropathicity were 0.669 (Table 3). The other detailed results of the physicochemical properties for the protein, including the number of amino acids, formula and isoelectric point were summarized in Table 3. |
| Prediction of signal peptide and transmembrane domain |
| We analyzed the signal peptides in NAT9 by using the SignalP 3.0 Server (http:// www.cbs.dtu.dk/ervices/SignalP/). Significant signal peptides were predicted based on the putative protein sequence, which indicated that NAT9 was secretory proteins and might play an important role (Table 4). The TMHMM Server v. 2.0 (http://www.cbs. dtu.dk/services/TMHMM/) were also used to analyze transmembrane domains. No transmembrane domains were found in this protein, which indicated thatNAT9 may locate in the cellular cytoplasm. |
| Prediction of secondary structure and construction of 3D structure model |
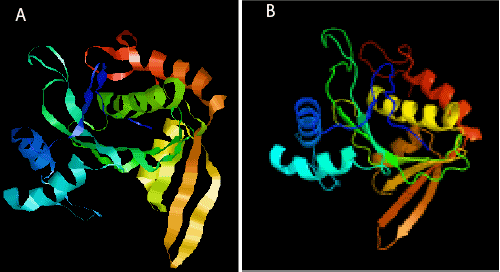
| The secondary structure of the putative protein was predicted by Predictprotein server software (http://www.predictprotein.org/). The percentage of α-helix, β-sheet, and random coil was 45.05%, 4.95%, and 50.00%, respectively. The 3D structures of the putative NAT9 protein were predicted by the SWISS-MODEL, using the GANT (Gcn5-related N-acetyltransferase) as the template. Disorder prediction and fold recognition showed that P. fulvidraco NAT9 protein was similar with c3tcvB_ genetic code template structure (Figure 4). Compared the template and P. fulvidraco NAT9 PDF files by Rasmol view fish; we found two monomer structure were very similar (Figure 4). Gcn5-related N-acetyltransferase super family is a comprehensive super family. They have a huge number of homologous region and acetylated related motif similar kind of material [8], including general control nonderepressible-5’, Hat1, Elp3, Hpa2, other eukaryotic and prokaryotic acetyltransferase of different substrates. The P. fulvidraco NAT9 code guarantees may be as an auxiliary activation factor with endogenous type A activity of HATs. |
| Discussion |
| In the current study, we firstly cloned the full-length cDNA of P. fulvidraco NAT9 gene. Through sequence analysis, it can be seen that the encoding protein of P. fulvidraco NAT9 gene is highly homologous with NAT9 in other fishes. This implied that the NAT9 genes were highly conserved in some fishes, and NAT9 of yellow catfish might have similar functions as in other fishes. NAT9 protein of yellow catfish has new feature compared with other fishes’ NAT9, hints that P. fulvidraco NAT9 gene has some unique functions than other fishes. |
| Tissue expression analysis shown yellow catfish NAT9 gene has huge differentially expressed in detected tissues. The suitable explanation for this under current conditions is that at the same time those biological activities related to the mRNA expression of yellow catfish NAT9 gene were presented diversely in different tissues. This indicates that NAT9 gene has diverse roles in different tissues because of the expression difference. To further understand the function of this novel gene, more research based on these primary results is needed. |
| The structure of P. fulvidraco NAT9 protein is similar to Gcn5- related N-acetyltransferases. Gcn5 have auxiliary activation factor with endogenous type activity of HATs [9]. NAT9 may be as an auxiliary activation factor with endogenous type A activity of histone acetyltransferases (HATs). HATs maintains cell dynamic equilibrium from histone acetylation/deacetylation [10]. The steady-state level of histone acetylation is a balance between the action of HATs and histone deacetylases. Obviously, as a result of histone acetylation plays an important role in the regulation of gene expression. So the unbalance of gene expression regulation will lead to cells disorder. It has been found that many diseases incidence relate to histone acetylation/deacetylation balance disorder. |
| In summary, based on the sequence from full-length cDNA of the NAT9 belong to NAT family in P. fulvidraco, we analyzed and detected tissue transcription profile and also predicted the putative protein using bioinformatics tools. These results of our study will pave the way to furthers researches the reproduction and disease resistance of NAT9 in P. fulvidraco. |
| Acknowledgment |
| The research was funded and supported by grants from National Science Foundation of China (31302177), and The authors would like to extend their sincere appreciation to the Deanship of Scientific Research at King Saud University for funding this Research No. RG 1435-012. |
References
|
Tables and Figures at a glance
| Table 1 | Table 2 | Table 3 | Table 4 |
Figures at a glance
 |
 |
 |
 |
| Figure 1 | Figure 2 | Figure 3 | Figure 4 |
Relevant Topics
- Analytical Biochemistry
- Applied Biochemistry
- Carbohydrate Biochemistry
- Cellular Biochemistry
- Clinical_Biochemistry
- Comparative Biochemistry
- Environmental Biochemistry
- Forensic Biochemistry
- Lipid Biochemistry
- Medical_Biochemistry
- Metabolomics
- Nutritional Biochemistry
- Pesticide Biochemistry
- Process Biochemistry
- Protein_Biochemistry
- Single-Cell Biochemistry
- Soil_Biochemistry
Recommended Journals
- Biosensor Journals
- Cellular Biology Journal
- Journal of Biochemistry and Microbial Toxicology
- Journal of Biochemistry and Cell Biology
- Journal of Biological and Medical Sciences
- Journal of Cell Biology & Immunology
- Journal of Cellular and Molecular Pharmacology
- Journal of Chemical Biology & Therapeutics
- Journal of Phytochemicistry And Biochemistry
Article Tools
Article Usage
- Total views: 9805
- [From(publication date):
March-2016 - Apr 03, 2025] - Breakdown by view type
- HTML page views : 8977
- PDF downloads : 828
