Review Article Open Access
Changing Paradigms in Cell Biology: Their Implication and Possible Applications
| Engwa Azeh Godwill* | |
| Chemical Sciences Department, Faculty of Natural and Applied Sciences, Godfrey Okoye University, P.M.B 01014 Thinkers Corner, Enugu Nigeria | |
| Corresponding Author : | Engwa Azeh Godwill Chemical Sciences Department, Faculty of Natural and Applied Sciences Godfrey Okoye University, P.M.B 01014 Thinkers Corner, Enugu Nigeria Tel: (+234) 8068473306 E-mail: engwagodwill@gmail.com |
| Received September 22, 2015; Accepted October 13, 2015; Published October 20, 2015 | |
| Citation: Godwill EA (2015) Changing Paradigms in Cell Biology: Their Implication and Possible Applications. Biochem Physiol 4:184. doi: 10.4172/2168-9652.1000184 | |
| Copyright: © 2015 Godwill EA. This is an open-access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited. | |
Visit for more related articles at Biochemistry & Physiology: Open Access
Abstract
The evolution of cell biology in the 20th century has evoked some remarkable changes in the existing paradigms. The discovery of alternative splicing, overlapping genes and protein splicing has shifted the one gene-one protein paradigm to several proteins. Proteins are no more the only catalytic macromolecule since the discovery of ribozymes. Genetic material was known to be stable in the genome and not mobile until the discovery of transposons. RNA editing has made possible the identification of certain missing genes and viruses are no longer considered the smallest infections agents after prions were discovered. Though not conclusive, recent findings suggest the possibility of transcription to occur in the nucleus. All these ground breaking discoveries are making substantial breakthrough in biological sciences particularly in understanding some cellular processes, their mechanisms on gene expression and regulation as well as the pathogenesis of certain illnesses. Also, these discoveries have revealed some potential drug targets and presently, some of the implicated biomolecules are being exploited for various pharmaceutical and biomedical applications.
| Keywords |
| Paradigm; Cell biology; Prion; Ribozyme; Alternative splicing; Nuclear translation; Overlapping genes; RNA editing; Transposon; Protein splicing; Biomedical application |
| Abbreviations |
| Ser: Serine; Cys: Cysteine; Asn: Asparagine; ADA: Adenosine Deaminase; LINE: Long Interspersed Nuclear Elements; SINE: Short Interspersed Nuclear Elements; LTR: Long Terminal Repeats; IS: Insertion Sequence; PrP: Normal Prion Protein; PrPC: Cellular Prion Protein; PrPSc: Scrapie or Pathogenic Prion Protein; mRNA: Messenger RNA; tRNA: TransferRNA; PTC: Premature Termination Codon; NMD: Nonsense Mediated Decay; SMN: Survival Motor Neuron; EIPA: 5:(N:Ethyl:Nisopropyl): Amiloride; U: Uracyl; VLDL: Very Low Density Lipoproteins; IDL: Intermediate Density Lipoproteins; LDL: Low Density Lipoproteins; A-to-I: Adenine to Inosine; HDV: Hepatitis Delta Virus; EST: Express Sequence Tag; CRISPRs: Clustered Regularly Interspaced Short Palindromic Repeats; Cas: CRISPR:Associated Proteins; HIV: Human Immunodeficiency Virus |
| Introduction |
| As time changes, so has new discoveries reshaped some concepts, theories and facts in cell biology. In the early nineteenth century, researcher had concluded on certain paradigms and theories in cell biology which were centered on the central dogma of life. Proteins were known to be the end product of gene expression without alteration in the transferred information. Thus, a single gene codes for one unique protein. This paradigm had been the basis for understanding cellular processes for various applications. It wasn’t too long for challenging and contrasting objections as a result of new molecular technologies to emerged and fine-tune the focus and interest of scientists. |
| Since the mid nineteenth century, scientific research especially in cell biology has tremendously grown with ground breaking discoveries which have now counteracted previous existing theories or paradigms. Little could someone in the early nineteenth century be expected to believe that proteins are not the only catalytic macromolecules [1], protein synthesis may occur in the nucleus [2] or viruses are not the smallest infective agents [3], to name a few. Since then, these non-existing paradigms are becoming more evident and realistic in life. There has been a change or shift in some existing paradigms in biology which are now being considered for new theoretical positions to understand and explain certain life processes in the cell and also for possible real life applications. The interest of this review is to present some of these ground breaking discoveries with their implication in cell biology and biomedicine and as well as their possible applications in the society. |
| Review |
| Prion |
| For a long time before its discovery, viruses were generally accepted as the smallest infective agents to cause a disease. Before the name prion was coined, a series of neurological diseases such as Creutzfeldt– Jakob disease in humans, transmissible spongiform encephalopathies” or Kuru, Bovine spongiform encephalopathy or mad cow disease, to name a few, triggered research to understand the etiology [4,5]. The early findings by Carleton Gajdusek which suggested that these infections were transmissible and had an infectious etiology rewarded him with the Nobel Prize for medicine in 1970 [5]. This agent was also thought to be a virus since they could pass through filters with a small pore size [6]. But later, this suggestion was rejected because infectivity was not affected by treatments that would usually inactivate nucleic acids such as ultraviolet light and nucleases [7]. In addition, this disease did not elicit immune response unlike viruses. By 1967, it was suggested to be a protein as a protease-resistant sialoglycoprotein was isolated from infected brain homogenates by Griffifh [8]. Later in 1982, Prusiner coined the term prion from its nature as a proteinaceous infectious particle to distinguish this infectious particle from viruses and designated the protein content of this Prion as PrP which fetched him a Nobel Prize [9]. Today, prions are now known to be the cause of these neurogenerative diseases that affect the brain. |
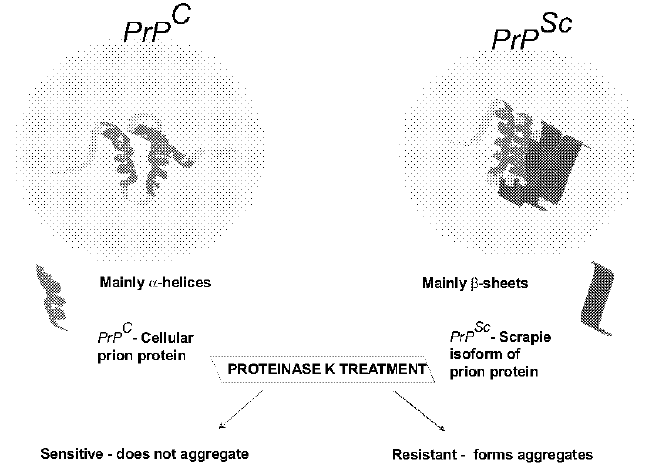
| A central biochemical feature of prion diseases known as the “prion hypothesis” is the conversion of normal prion protein (PrPC) to an abnormal, misfolded, pathogenic isoform designated PrPSc, (named after “scrapie,” the prototypic prion disease) [10]. A chromosomal gene encodes PrPC (the cellular isoform of PrP) and no PrP genes are found in purified preparations of prions [11]. The normal form of the protein, referred to as PrPC, is a highly conserved cell surface protein attached via a glycophosphatidyl inositol anchor. It is expressed in a wide range of cell types, and particularly in neuronal cells. PrPC is a sialoglycoprotein of molecular weight 33 to 35 kDa, with a high content of α-helical secondary structure that is sensitive to protease treatment and soluble in detergents. The disease associated isoform, referred to as PrPSc, is found only in infected brains as aggregated material, is partially resistant to protease treatment and insoluble in detergents, and has a high content of β-sheet secondary structure. PrPSc is derived from PrPC by a posttranslational process whereby PrPSc acquires high betasheet content and a resistance to inactivation by normal disinfection processes. The PrPSc is less soluble in aqueous buffers and when incubated with protease (proteinase K), the PrPC is completely digested (sometimes indicated by the “sensitive” superscript, PrPsen) while PrPSc is resistant to protease (PrPres). Unlike cellular form (PrPC) that does not aggregate, the pathogenic form (PrPSc) aggregates together (Figure 1). |
| Prions do not contain nucleic acid and so a big question was how do they replicate or multiply? The only possible hypothesis which has not been justified is the only-protein hypothesis postulated by Prusiner, who proposed a model of infection that relied on a misfolded “scrapie” form of the normal prion protein molecule (PrPSc) being able to induce the refolding of the host’s constitutive cellular prion protein (PrPC) to mimic its aberrant conformation [12]. This proteinaceous infectious particle (prion) would encode differences in disease pathogenesis by differences in its conformation, allowing for the transmission of varying “phenotypes”. This idea was supported by Kellings and colleagues who demonstrated that no group of similar DNA fragments consistently copurified with PrPSc [13]. |
| The discovery of prion has brought new insight in the etiology of some neurodegenerative diseases and has now provided new prospects on possible targets for drug discovery and management of this category of diseases. More so, the postulation of the only-protein hypothesis which suggests that proteins can induce the refolding of another protein without the influence of a genetic or external factor has led to a new paradigm of “protein to protein conversion”. This paradigm is now postulating new ideas in cell biology. If a protein can influence the structure or conformation of another, PrPC may be of great therapeutic interest to certain diseases which may be due to misfolding of proteins such as self-antigens or other similar diseases of protein origin. Also, due to the ability to influence refolding of proteins and their very small size, prions can possibly be exploited as gene therapies for site-specific treatments. This can be achieved by influencing a change in their conformation through alteration of the pH and temperature to serve as nanomolecules in carrying drugs to various target sites. |
| Alternative splicing |
| Naturally, the central dogma of life is centered between DNA, RNA and protein in this order. DNA the main nucleic acid that codes for protein necessitates the presence of RNA to achieve this transformation. A gene in DNA is first transcribed to RNA, particularly mRNA, which is then translated to a protein. However, before an mRNA can do this, it needs to be matured. Maturation of mRNA requires a mechanism to convert the pre-mRNA formed after transcription. In this process, the non-coding sequences called intron of the pre-mRNA is excised leaving behind only the coding sequences (exons) which are joined together to become a matured mRNA. Splicing as the name implies require some enzymes to cut or splice off some sequences and join others. The goal of mRNA splicing is to produce mature mRNA transcript that will translocate to the cytoplasm where it is converted to a protein. Because only one transcript can be formed from a gene and all things being equal, the concept of one gene to one transcript holds. This had been the existing paradox until the human genome was sequenced [14]. |
| Scientists thought that the complex DNA of a human was made up of as many as 150,000 different genes based on the number of different transcripts (mRNAs) which had been found in humans assuming that there should be one gene for each mRNA. However, this became controversial when the human genome was sequenced and found it to contain only about 32,000 genes or even less [15]. Scientists started to question the paradox of one gene-one transcript and suggested the possibility of many transcripts for a gene. To support this claim, they went further to do proteomics analysis as well as sequence various proteins and aligned them for homology [16]. This was the basis of a new paradigm of one gene and many proteins. Further research on mRNA splicing suggested that many transcripts could result from a single gene by varying mechanism of splicing which they later called alternative splicing [17]. |
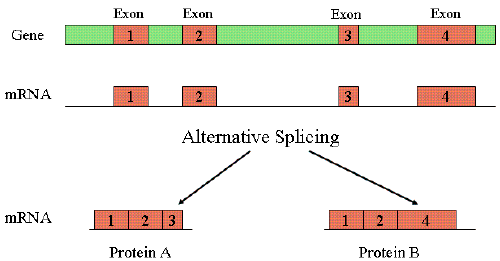
| Alternative splicing is a process which allows the production of a variety of different proteins by generating several transcripts from a single gene only. Since most genes in eukaryotic genomes consist of so many exons and introns, it is accomplish through splicing by removing introns from the pre-mRNA and joining several exons ends together in a random manner such that based on the type of information to be conveyed, several different types of exon combinations can be done to give rise to many different transcripts of a single gene which in turn gives rise to diverse proteins (Figure 2) [18]. In humans, it is estimated that alternative splicing occurs in more than 60% of genes [19]. |
| Even though understanding the mechanism that governs alternative splicing has been very challenging, it has brought new insight to processes of gene expression, regulation and cause of certain diseases [20,21]. Through alternative splicing, gene expression can be regulated positively and negatively. In the case of positive regulation, the splice pattern could be such that it selects the desired exons for a requested protein. For a negative regulation, it could be such that it brings together “non-compatible” exons persee, thereby generating a wrong reading frame leading to a nonsense mediated demolition (NMD) or pathology. |
| Alternative splicing has been implicated in a large number of human pathologies such as neurodegenerative [22], cardiovascular [23], cancer [24] and some other diseases but with promising targets for therapeutic intervention [25]. A therapeutic target could be such that, for a disease with a defect of an excluded exon(s) in a functional protein, certain key exons implicated are included in the transcript. For example, in survival motor neuron (SMN) diseases, certain small drug molecules such as sodium vanadate, aclarubicin, indoprofen, hydroxyurea, valproate, 5-(N-ethyl-Nisopropyl)- amiloride (EIPA), and phenylbutyrate that increases inclusion of exon 7 of SMN2 gene have been identified [26]. Some targets could be to increase or enhance the expression of certain transcripts in diseases which are as a result of deficiencies. Novantrone is one of such drug that can enhance the effectiveness of therapeutic treatments for familial neurodegenerative diseases by stabilizing the tau pre-mRNA splicing regulatory element [27]. Another therapeutic target is by suppressing alternative splicing in cases where it favors the expressions of proteins implicated in disease conditions. For example, Meayamycin, is active against multidrug-resistant cells and performs antiproliferative effect against human breast cancer MCF-7 cells by suppression of alternative splicing [28]. |
| Though alterative splicing is promising for new therapeutic solutions, effective treatment requires accurate diagnosis and identification. This has been one of the major challenges in the management of illnesses which are as a result of alterations in gene expression mechanisms. Splicing regulators which are key markers of such diseases usually exists in combination to define a diseases state. These associative components need to be clearly and specifically identified in their various cascade combinations to indicate a particular illness or disease trait. Protein microarray diagnostic approach may be of future prospects to identify splice regulators involved in such illnesses for effective and accurate diagnosis to ensure more specific and reliable treatment. |
| RNA editing |
| As previously described above, the central dogma of biology stated that genetic information flows from DNA↔RNA→Protein. This implies that amino acid sequence of a protein will directly reflect the genetic code of the nucleic acid sequence of the transcribed gene. In other words, the nucleic acid sequence of mRNA is a direct copy of the sequence of DNA. This had been the existing paradigm but became questionable when this paradigm was not binding for all genes. In certain organisms, some protein sequences did not show a complete reflection of the nucleic acid sequence. The first challenge to this idea led to the discovery of intervening sequences within genes of higher organisms which are precisely spliced out of the mRNA, and the coding RNA fragments, or exons, are then joined together to create the complete gene. More so, further research came up with evidence obtained with an ancient group of parasitic flagellated protozoa, the kinetoplastids, showed that the sequence of nucleotides in mRNAs in coding regions can be modified after transcription such that the sequence of the protein obtained was altered [29,30]. |
| The phenomenon known as RNA editing was used to describe such deviations from the existing paradigm. The first evidence of RNA editing came up when Benne and colleagues in 1986 found 4 extra Uracyls (U’s) in the conserved frameshift region of the COII mRNA sequence that were not encoded in the maxicircle DNA of Trypanosomia Brucei [31]. Also, another study with Leishmania tarentolae showed that the Cyb mRNA was edited within the 5’ end by the insertion of 39 U’s at 15 sites, thereby creating 20 new amino acids at the amino end of the protein, including an AUG that encode methionine for initiation of translation [32]. Apart from addition, deletions of U’s were also found to occur in some genes, such as the COIII gene of L. tarentolae, although at a lower frequency [33]. Cryptogene (hidden gene) was the word coined to describe genes whose transcripts are edited within coding regions. Also, pre-edited region was the term used to describe the region of an mRNA which is to be edited [34]. Pan editing was a term used to describe an extensive editing with addition of over hundred U’s into the mRNA. |
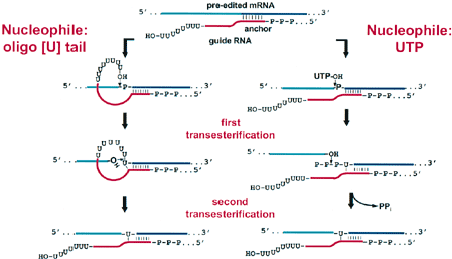
| Another important insight of RNA editing came from the findings of Abraham and colleagues in 1988 who found out that the pan-editing of the COIII mRNA of T. brucei appeared to occur in an overall 3’ to 5’ direction. This finding suggested that RNA editing was done only after transcription since it moves in a 5’ to 3’ direction [35]. Subsequently, the process on how RNA editing occurs was revealed following the discovery of certain short RNA sequences called guide RNA (gRNA) which contained sequences at their 5’ end that could base pair with the mRNAs just downstream of the pre-edited regions [36]. Later, certain enzymes were discovered to be responsible for the transfer of U’s across the bound RNA molecules. The first was a terminal uridylyl transferase or TUTase responsible for the addition of U’s to the 3’ end of the gRNAs [37]. Also, a mitochondrial RNA ligase was shown to covalently link together two RNA molecules [37]. Since then, a series of enzymes and protein factors have been implicated in the process of RNA editing (Figure 3). |
| One major achievement of RNA editing is that it made known the discovery of certain missing genes (cryptogenes) whose transcript could not be identified. For example, Feagin in 1988 discovered the missing COIH gene in T. brucei which was actually present throughout, but was a truly hidden cryptogene since the transcript was so extensively edited with hundreds of U additions over almost the entire length that the mature edited mRNA was nearly twice the size of the gene [38]. Also, RNA editing has exposed new therapeutic targets for diseases. One of such target is a variant of Apolipoprotein B. Apolipoproteins are essential components of plasma lipoproteins that serve as transport vehicles of lipid nutrients in the circulation [39]. The unedited apolipoprotein B transcript gives rise to full-length apoB (apoB100), which is the major protein component of very low density lipoproteins (VLDL) and their maturation products intermediate density lipoproteins (IDL) and low density lipoproteins (LDL) and has been shown to increase the susceptibility to atherosclerosis. On the contrary, a truncated apolipoprotein B (apoB48) which has undergone tissue-specific base modification editing showed much less atherogenic potential [40]. Understanding the RNA editing mechanism and its regulation might eventually be used therapeutically to decrease the risk of atherosclerosis in humans. Also, an Adenine to Inosine (A-to-I) editing within the antigenomic RNA of the subviral human pathogen Hepatitis Delta Virus (HDV), has been shown to repress replication [41]. This could also be a potential target for anti-replication drug for the virus. |
| More so, recent research on RNA editing has led to the discovery of powerful target specific regulatory systems known as the clustered regularly interspaced short palindromic repeats (CRISPRs) and CRISPR-associated (Cas) protein [42]. These systems exploit guide RNA to specifically identify and eliminate foreign nucleic acids introduced by invading phages and conjugative plasmids there by creating memory in the cell for subsequent infections. A new paradigm of adaptive molecular immunity has emerged to explain this discovery. This new technology is regarded as one of the most powerful biotechnological advances of our time. However, so much is being done to ensure that the regulatory system specifically identify only target nucleic acids. If such system effectively works, it may be of importance to exploit it for other viral molecules such as HIV which have been very difficult to eliminate as it evade the natural adaptive immunity in humans. |
| Overlapping genes |
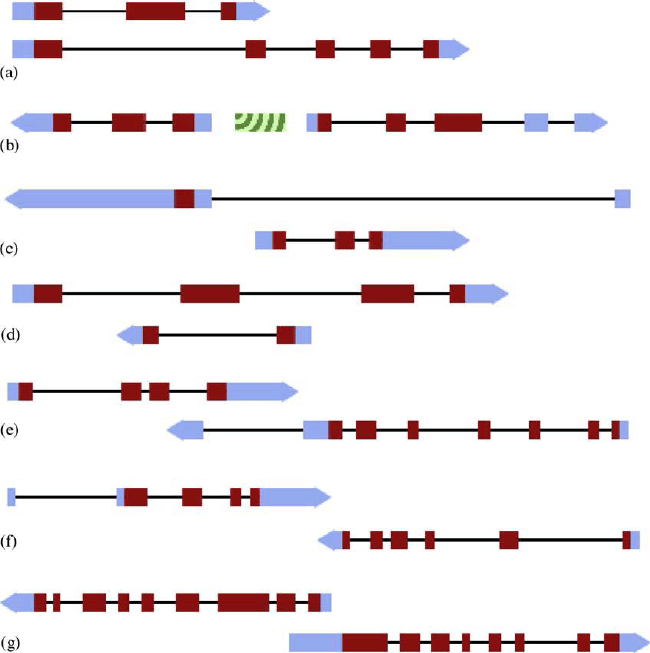
| Normally by principle, a codon is composed of three nucleotides which codes for an amino acid. A gene is usually defined by a start codon which initiates the transcription start site. With only one start site, a transcription unit is known to produce only one transcript, hence a single polypeptide. However, since there are 3 bases for a codon, there are 3 possible reading frames of codon in the same gene. Occasionally, it is possible for the same stretch of genome codes for more than one protein in different reading frames. This wasn’t the case until Barrel and colleagues in 1976, showed the possibility of several proteins for a transcript by their discovery of overlapping genes in bacteriophage Ф X174 [43]. This was another deviation from the one gene-one protein paradigm. Overlapping genes generally refer to pairs of genes that overlap in their transcribed sequences. After this discovery, it took another decade before similar observations were noticed in higher eukaryotes. In 1986, overlapping genes were identified in Drosophilia [44] and mouse. Since then, several overlapping genes have been discovered in humans by large scale expressed sequence tag (EST) and genome sequence studies [45,46]. The overlapping genes discovered could be classified in different categories [47]. Firstly, it could be such that different reading frames of a gene could result to two or more proteins. In this type of overlap, transcripts share the same locus on the same DNA strand. Another type of overlapping genes is those on opposite strands sharing the same promoter region. In such case, the promoter functions bi-directionally. Here, though genes are located on opposite strands, the overlap is only in the promoter region and transcripts do not share any other sequence. This overlapping could lead to a sense transcript when it is in the forward direction of the promoter or an antisense transcript when it is in the reverse direction of the promoter. Another category of overlapping genes is those nested within another gene. That is, they are found within the transcript of a large gene. This could also be in the intron of a gene which can be transcribed (Figure 4). |
| Antisense transcripts have been thought to play major roles in regulating gene expression [47]. Antisense transcripts can bind with the sense transcript to inhibit its transcription by a process known as transcriptional interference. RNA masking is another mechanism of gene regulation in which the antisense mRNA can bind the sense mRNA thereby inhibiting the alternative splicing machinery by blocking the accessibility of regulatory factors. Also, the double-stranded RNA (dsRNA)-dependent mechanisms is possible in which binding of the sense and antisense transcript can lead to gene silencing by RNA editing, or RNA interference (RNAi)-dependent gene silencing. Large doublestranded regions may be modified by the RNA-editing machinery. |
| Due to the ability of antisense transcripts to alter gene expression, they are thought to be implicated in so many human disease conditions [48]. The proliferation of endometrial cells observed in patients with endometriosis, is thought to be due to reduced expression of the exon 1B isoform of the basic fibroblast growth factor (bFGF) antisense transcript [49]. Tenascin X gene, which overlaps with last exon of the CYP21 gene, is essential in regulation of collagen deposition by dermal fibroblasts and has a causative role in human Ehlers-Danlos syndrome [50]. Human PRG4 gene overlapping at 3’ end with TPR gene is involved in the development of arthropathy-camptodactylly syndrome [51]. |
| In as much as it is known that overlapping genes are involved in gene regulation, which a positive regulation will be helpful to suppress, certain illnesses in humans, negative regulation are usually consequential. Overlapping genes in microorganisms is one of the major ways that microbes evade host immune system by alternating their gene expression to prevent the host from eliciting specific immune responses. It becomes primordial to clearly understand overlapping genes and the expression pattern in such pathogenic microorganisms for potential therapeutic targets. |
| Ribozyme and artificial DNA enzymes |
| Enzymes are generally biochemical catalyst that speeds up reaction either by building or breaking down (cutting) molecules. Ever since the discovery, it was concluded that only proteins can possess this activity. This theory was never challenged until DNA and RNA were sequenced and scientist observed that RNA molecules could be shortened after transcription. This notion was glaring and unclear to the minds of many till the 1980s when the works of Thomas Cech and Sidney Altman made a breakthrough on the concept of RNA to have a double role; not just for the synthesis of proteins from DNA but also, an intrinsic catalytic role [52]. The findings of this notions of catalytic RNA was shown by Thomas Cech and his group at University of Collorado in 1982 when they found that an RNA in the ciliated protozoan Tetrahymena thermophila could cleave and splice itself without any external protein or energy source [53]. Similar findings by Altman’s group at Yale University supported this claim when they found an independent RNA as an enzyme called ribonuclease P in Escherichia coli which was able to process transfer RNA (tRNA) precursors without any protein factors [54]. From this concept that RNA has catalytic activity, the name ribozyme was coined to describe such catalytic RNAs. Thanks to this discovery, Cech and Altman shared the noble prize for chemistry in 1989 as a reward for their contribution to knowledge in science. Knowing that certain RNA could catalyze reaction, Cech made another discovery to explain how transcribed RNA could be shortened. Cech identified a non-coding region of RNA which undergo self-catalysis and called it intron. Since then, some introns have even been shown to contain sequences that can encode proteins required for the processing of DNA and RNA. |
| Following the discovery of ribozymes, it is now being claimed that RNA was the first product of the prebiotic soup. In other words, RNA was the first macromolecules to have emerged before DNA and protein. This notion described as the RNA world hypothesis stated that DNA and protein-based life was preceded by RNA based life, in which RNA acted as both the genetic material and cellular enzyme [55]. As cellular metabolism became more sophisticated, increasing demands created the transition to protein-based enzymes and more stable genetic information (in the form of DNA). |
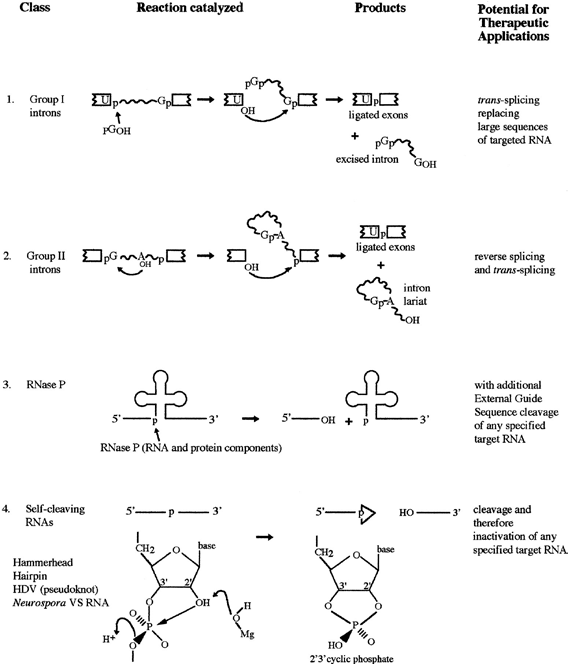
| Based on the characteristic structure and reaction mechanism, ribozymes have been distinguished into various classes [56]. The two major classes of ribozymes are distinguished either by self-splicing or selfcleaving. The first class consist of self-splicing group I and II introns and the Ribonuclease P while the second class consist of the hammerhead, hairpin, hepatitis delta ribozymes and varkud satellite RNA which are self-.cleaving ribozymes. Group I and II ribozymes generally excise themselves from RNA molecules and require co-factors. The distinguishing factor between them is that group I require guanosine in addition to other metallic co-factors while group II do not. Ribonuclease P is a ribonucleoprotein consisting of approximately 375-nucleotide RNA plus a small polypeptide whose catalytic activity lies in its RNA subunit. The RNA portion cleaves tRNA precursors to produce the mature tRNA. The second class of ribozyme generally functions as self-cleavers. They are usually microsatellite RNAs that cleave themselves from precursor RNA without their involvement in splicing (Figure 5). Size is another factor used to distinguish the catalytic mechanism of ribozymes. Large ribozymes of several hundred up to 3000 nucleotides generate reaction products with a free 3’-hydroxyl and 5’-phosphate group. In contrast, small catalytically active nucleic acids which range from 30 to 150 nucleotides in length generate products with a 2’-3’-cyclic phosphate and a 5’-hydroxyl group. |
| With catalytic activity, capability of cleaving mRNA molecules in a specific sequence and the selective ligation with target mRNAs, ribozymes can be specifically tailored for the suppression of particular genes [57]. Group I intron ribozyme can be specifically designed to repair abnormal mRNA molecules and has been shown to be suitable for the correction of deficient mRNAs [58]. For example, a trans-active group I intron was used to repair mutant β-globin RNA in erythrocyte precursors from patients with sickle cell anemia by replacing the mutated part of the β-globin RNA by the γ-globin-3’-exon. In addition, attempts have been reported to address diseases caused by trinucleotide repeat expansions, including Huntington’s disease and myotonic dystrophy [59]. Another therapeutic class of interest is the group II introns. It has been demonstrated that group II introns can be redirected to insert themselves into therapeutically relevant DNA target sites in human cells [59]. This is due to the fact that they are able to insert themselves into an intron less allele on the DNA level by reverse splicing and reverse transcription, a process called retro homing. |
| Ribozymes have also been targeted as anticancer agents especially at genes involved in signal transduction cascades, such as genes for expression of growth factors and their corresponding receptors, genes for the induction or progression of tumors. Also, genes of tumor angiogenesis and the genes important in cancer therapy such as MDR- 1-multidrug resistance are potential targets. Two clinical trials for cancers are ongoing to evaluate the potentials of therapeutic ribozymes; Angiozyme are being examined in a phase II trial for treatment of metastatic colorectal cancer [60] and herzyme in phase I clinical trials to determine toxicity and efficacy in breast and ovarian cancer patients [61]. |
| Ribozymes can be used to study the function, regulation and expression of genes. They provide a unique tool for understanding gene function because they allow one to assess cellular responses to a rapid ablation of target gene expression. They are also unique in that they can inactivate specific gene expression, and thereby can be used to help identify the function of a protein or the role of a gene in a functional cascade. The use of ribozymes for target validation is critical for both basic biological research and drug discovery. Compared to other means of target validation such as use of transgenic animals, ribozymes offer specificity and ease of design and usage [62]. |
| Another function of ribozyme is their possible use as biosensor to detect analytes. Biosensors are designed such that catalytic activity is regulated by binding of a small-molecule ligand remote to the catalytic site in a modular fashion. Many artificial allosteric ribozymes have been identified, such as several hammerhead ribozyme variants for such activity [63]. Interestingly, allosteric ribozymes have shown to response to more than one ligand since their catalysis is regulated by oligonucleotide subunits [64]. |
| The fact that DNA is structurally similar to RNA, the possibility of them having potential catalytic activity has also been questioned. Till date, no natural DNA enzyme has been identified. However, the word deoxibozymes was brought to existence by Breaker and Joyce who showed artificial DNA to possess catalytic activity in vitro [65]. This was achieved through an in vitro selection technique which was applied to large populations of random-sequence DNAs, leading to the recovery of specific DNA enzyme that catalyzes the Pb2+-dependent cleavage of an RNA phosphoester in a reaction that proceeds with rapid turnover whose catalytic rate was comparable to that of known RNA enzymes. Since then, synthetic deoxiribozymes have been exploited for their function as biosensors. For example, a deoxyribozyme sensor for metal ions and small organic molecules whose sensing ability is based on fluorescence or colorimetric signals has been developed [66]. With such developments, it will be no news in future if natural occurring deoxiribozymes are discovered. |
| As a new class of pharmaceutically important compounds, ribozymes and deoxiribozymes offer promising potential in the treatment of certain diseases and genetic disorders. However, the major concern is how to specifically deliver these therapeutic molecules to their target site. This entails the development of safe, effective and tissue specific delivery systems. Both viral and non-viral vectors have been exploited as delivery systems [67], but because viral DNA can easily integrate into host genome and be expressed, viral vectors offer a promising potential for cell specific delivery of ribozymes. If such delivery approaches along with more precisely targeted expression systems are developed, there is future hope to have novel pharmaceutical agents for the treatment of some diseases. |
| Transposons |
| Before her discovery, genetic materials were known to be stable and intact within the genome. However, there was no doubt that the functions of certain genes could be altered by mutation. Generally, mutants are environmental materials foreign to the genome that can cause changes in base sequences either by base addition, deletion or change. Mutation could therefore lead to genetic variation between species or organisms but was not by self. Also, during cell division in sexual reproduction, chromosomes divide and there is homologous recombination which permits the exchange of genetic material between related organisms. However, genetic material could only be exchanged between loci of similar genetic material (alleles). Even though this recombination leads to genetic diversity, after division, the recombined chromosomes are similar and stable without any breakages. |
| Strangely around 1940, Barbara McClintock, a geneticist while studying the color of maize (Zea mays) kernels by microscopic observation of chromosomes, found an unusual breakage of chromosome 9 in one maize strain very frequently at a particular locus [68]. She postulated that this breakage was due to a genetic element as she identified two genetic elements; one at the site of the breakage called Ds (dissociation) which she believe to have caused the breakage and the other Ac (Activator) which is required to activate the breakage [69]. McClintock began to suspect that Ac and Ds were actually mobile genetic elements when she found it impossible to map Ac to a particular position. In some plants, it mapped to one position; in other plants of the same line, it mapped to different positions. She also observed that rare kernels with dramatically different phenotypes could be derived from the original strain that had frequent breaks in chromosome 9. One of such phenotype was a rare colorless kernel containing pigmented spots. In trying to explain this phenomenon, McClintock even strongly supported her claim to describe these colorless kernels with spots as unstable phenotypes due to some mobile genetic elements which could cause the genome to be unstable. Even though McClintock postulate for transposable genetic element was accepted by geneticists, it was thought to be a rare situation and reluctant to consider the possibility in other organisms. The breakthrough came about 20 years later, when a new class of mutations in genes of a laboratory strain of the common intestinal bacterium Escherichia coli was found [70] and her postulate was later confirmed when the Ac and Ds elements were isolated by Fedoroff and colleagues in 1983[71]. |
| After E. coli, transposable elements were subsequently isolated from the genomes of many organisms, including Drosophila and yeast, it became apparent that Mcclintock postulate of transposable elements was true and was now considered a significant component of the genome of most and perhaps all organisms. In recognition of her work, Barbara McClintock was awarded the Nobel Prize in Medicine in 1983. |
| Since then, transposable elements have been identified in most living organisms. In prokaryotes, the first discovery was made in E. coli in the gal-operon [70]. Two types of transposable elements were discovered; the insertion sequence (IS) element and transposons. The IS elements are short mobile DNA sequence that do not carry genes other than those needed for their movement. They encode a protein, called a transposase, which is an enzyme required for the movement of IS elements from one site in the chromosome to another. In addition, all IS elements begin and end with short inverted repeat sequences that are required for their mobility. Transposons on the other hand are usually long, containing several genes and are flanked at both ends by IS elements. |
| In eukaryotes after the discovery in maize, transposable elements were subsequently discovered in Drosophilia (P element) [72]. DNA transposons were discovered in eukaryotes and found to show similar activities like the simple transposable elements in prokaryote. Also in eukaryotes, certain transposable elements were found to have similar activity like retroviruses [73]. This class was name retrotransposons. Just like retroviruses, these DNA elements are first converted to RNA intermediate, which after being spliced; the transposable fragment is later converted to DNA by reverse transcriptase which then integrates into another location in the genome again. Some of these elements contain long terminal repeats (LTR) and are called LTRretrotransposons. |
| In humans, the retrotransposons are of two types: long interspersed nuclear elements (LINEs) and short interspersed nuclear elements (SINEs) [74]. LINEs move like a retrotransposon with the help of an element-encoded reverse transcriptase but lack some structural features of retrovirus-like elements, including LTRs. SINEs can be best described as non-autonomous LINEs, because they have the structural features of LINEs but do not encode their own reverse transcriptase. Presumably, they are mobilized by reverse transcriptase enzymes that are encoded by LINEs residing in the genome. There are more than 1 million SINES in humans and are called Alu because it contains a target site for the Alu restriction enzyme. |
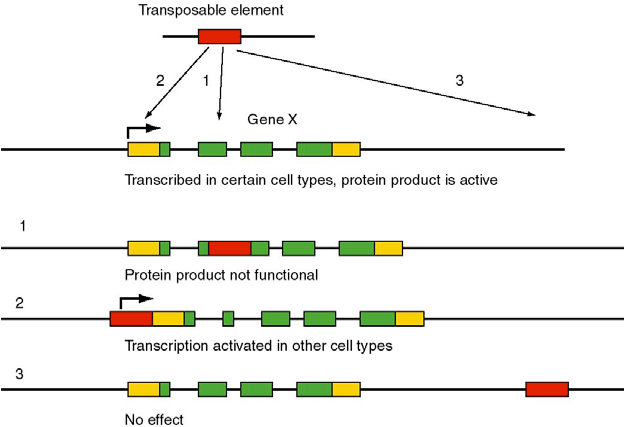
| Transposable elements which can freely move within the genome may have various implications. In situations where the mobile element inserts itself within a gene or an exon of a gene, that gene may be disrupted and becomes nonfunctional or transcribe a nonfunctional protein. In certain situations where it integrates at a regulatory site of a gene such as the promoter site, it may enhance transcription. Certain cases where transposable elements integrate at non coding sites or outside a gene, they may not have any effect on gene expression. Based on the above possibilities, transposable elements can be exploited for various genetic, biomedical and research applications (Figure 6). |
| Transposons are now becoming useful tools for genetic research in the discovery of new genes. They have been exploited to tag genes for cloning and to insert transgenes. One of the most exploited is the P element in Drosophila [75]. It provides one of the best examples of how geneticists exploit the properties of transposable elements in eukaryotes. P elements can be used to create mutations by inserting into genes, to mark the position of genes, and to facilitate the cloning of genes. The inserted P element into genes is identified using a probe. In such experiments, new genes or other genetic properties can be identified. Another important function of transposon could be their usefulness as gene therapies in the treatment of certain genetic linked diseases. Their ability to insert into specific sites in a genome, gives it the characteristic property of a gene therapy [76]. Because retrotransposons are similar to retroviruses, it is our hope that possible transposon gene therapies could be developed especially to cure genetic disorders. |
| Despite the numerous potential benefits, transposons are believed to be the cause of some viral disease. Because the sequence of LINEs resembles that of retroviruses, it is thought that certain viral diseases may be as a result of the displacement of LINEs within the genome to a locus of their promoter site where they can be transcribed. |
| Protein splicing and inteins |
| After the discovery of RNA splicing, it was widely concluded that the end product of a transcript was a fully matured protein which could undergo certain post-translational modifications but not splicing. This was farfetched until around 1990 where two groups found that a section of Saccharomyces cerevisiae Sce VMA1 vacuolar ATPase gene was absent in the mature ATPase protein [77,78]. The term protein splicing was born to explain this phenomenon. The discovery of protein splicing has help to explain some mechanisms which organisms utilize to enhance the function of some protein or rendering certain inactive to active enzymes. They predicted that an internal section of this protein was removed by protein splicing instead of RNA splicing and that a single gene encoded two stable proteins: the host protein (extein) and the intervening protein (intein) [79]. This idea was challenged by several researches to show that the phenomenon was due to RNA splicing but failed to demonstrate this in several attempts. This discovery was finally established when another group was able to clone and express a Pyrococcus species DNA polymerase intein between two unrelated proteins, resulting in temperature-dependent splicing [80]. This experiment did not only establish protein splicing mechanism but also led to another great discovery of inteins, known as homing endonuclease which was responsible for splicing of proteins. Homing endonucleases belonging to group I introns or inteins are a large class of site specific DNases, which are encoded by mobile genetic elements. |
| Since then, more than 200 inteins have been identified in living organisms [81]. The inteins range from 128 to 1650 amino acids and share a set of highly conserved sequence motifs. Majority of inteins identified are bifunctional in nature. They contain the endonucleases motif to cleave specific sites in the protein, as well as the characteristic motifs of a homing endonuclease that confers genetic mobility upon the intein encoding gene. Many inteins have been shown to self-splice in vitro without the requirement of external energy or protein cofactors [82]. |
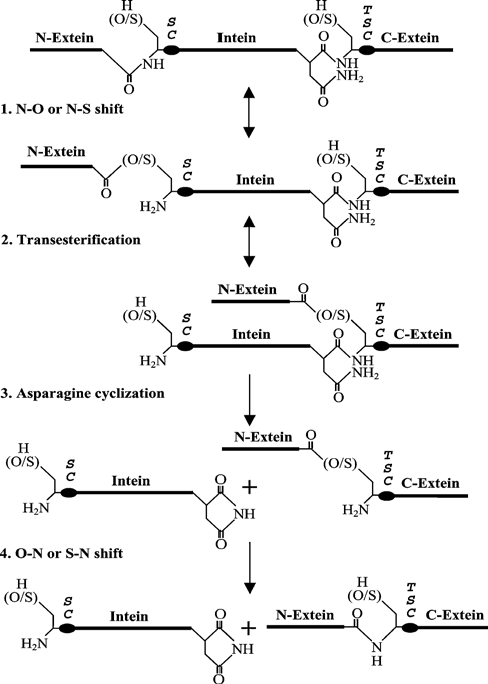
| Based on sequence signatures and splicing mechanisms, there are three classes of inteins [83,84]. The Standard class I intein splicing mechanism consists of the following; firstly, an acyl rearrangement to convert the N-terminal splice site peptide bond from an amide to a (thio) ester. Secondly, splicing is by trans-esterification to form a branched intermediate. Thirdly, Asn cyclization resolving the branched intermediate by cleaving the C-terminal splice site, and finally, a second acyl shift to form an amide bond between the ligated extein segments. Both class 2 and class 3 inteins can still splice, although they lack a Ser1 or Cys1 nucleophile and are thus unable to form the linear (thio) ester intermediate. Another class of inteins are naturally occurring transsplicing inteins in which a host gene is split into two separate coding regions, each fused to either the N-terminal or C-terminal portion of an intein-coding region [85].The full-length host protein is formed when the N-terminal and C-terminal intein regions come together to reconstitute protein splicing activity (Figure 7). |
| Since the discovery of inteins, it has been exploited for various applications. Intein-based methods by protein cyclization can be used to modify the sequence or structure of recombinant proteins [86]. Cyclization results in enhanced stability and bioactivity of the target proteins [87]. Also, cyclization has been used for in vivo generation of large libraries of genetically-encoded cyclic peptides for high throughput screens [88]. Intein technology can be used to control the toxicity of certain organism. This can be done by producing a nontoxic protein precursor by inserting an intein in a toxic protein [89]. Also, expression of antibodies using a single open reading frame was achieved by fusing the genes for antibody heavy and light chains with an intein [90]. This fusion protein was successfully expressed and processed in mammalian cells, with intein-directed N- and C- terminal cleavage reactions resulting in antibodies with the correct sequences for both heavy and light chains. |
| More so, inteins can facilitate in vivo gene modification by serving as genetic markers [91]. Muller and coworkers interrupted the Pch PRP8 intein with selectable markers, including aminoglycoside phosphotransferase and imidazoleglycerol-phosphate dehydratase. The interrupted inteins are able to splice, and could serve as selectable markers for expression of the spliced extein. They can also serve as biosensor for sensing protein-protein interaction, permit DNA methylation, protein localization and internalization, small molecules, protease activity, oxidation state etc. and facilitate segmental isotopic labeling in vivo, as well as the in vivo addition of chemical probes to specific target proteins. Intein has helped to facilitate the transfer of genes in other organisms (transgenic activity). Protein trans-splicing in mammalian cells and in mice has been used to test delivery of transgenes by adenovirus delivery vectors [92]. The findings suggest that inteins could be used for the in vivo generation of proteins too large to be delivered by traditional viral vectors. |
| Though protein splicing and intein mechanism of action has extensively been exploited for various applications, certain aspects need to pay considerable attention. The relation of inteins with regulatory proteins, their evolutionary trait and preference for some protein family’s needs to be addressed to better enhance their understanding. |
| Nuclear translation |
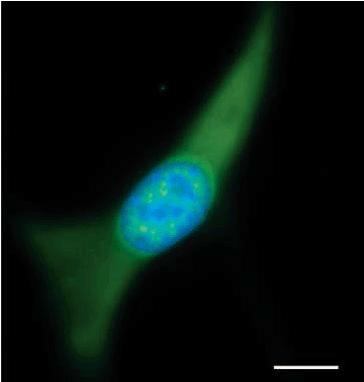
| In eukaryotes with defined nucleus, it has been known for years that protein synthesis is compartmentalized in the cytoplasm unlike in prokaryotes where the transcription and translation mechanisms are coupled together [93]. Because ribosomes are the main machinery for the synthesis of proteins and are localized in the cytoplasm, it was generally accepted that protein synthesis can only occur in the cytoplasm. Little did we believe that protein synthesis could occur in the nucleus (nuclear translation) but about forty years ago, some evidence emerged [94] even though not very convincing. The most convincing evidence was the findings of Cook and colleagues in 2001 that showed isolated nuclei to incorporate radiolabelled amino acids into nascent peptides [95]. Since then, so much concern has been laid to address this concept. Based on the findings made, there has been direct and indirect evidence in support of nuclear translation. Firstly, some fraction of nonsense mediated decay (NMD) was shown to occur in the nucleus [96,97]. Due to the fact that translating ribosomes are the only known means of detecting termination codons, and some NMD has been shown to occur within the nuclear fraction, it is believed that the NMD scanning mechanism utilizes active nuclear ribosomes [98]. Secondly, a study by Iborra and colleagues in 2004 showed the presence of nascent (newly synthesized) peptides (9-15%) to be found in the nucleus using biotinlabeling suggesting some occurrence of translation in the nucleus [99]. Thirdly, some elements of translation machinery such as ribosomal RNA, translation protein factors (IF2/eIF5B, eIF2α, eIF4E, eIF4G, eRF3etc.) have been shown to be present in the nucleus [100,101] (Figure 8). Even though this seems to be the case, arguments against nuclear translation have been raised focusing on the potential limits and strength of control of the supporting works. Other researchers have argued that the nuclear signal reported is a consequence of cytoplasmic contamination [102]. Another argument advanced is that over-permeabilization might lead to entry of cytoplasmic ribosomes into the nucleus, which then generates the nuclear signal [102]. However, though these arguments, their propositions have not completely counteracted the fact that to some extent, nuclear translation may occur as shown by the supporting findings. |
| If for any reasons the world assumes the position of advocates for nuclear translation, then it will be considered that the overall interest of the cell for this idea is firstly to control the quality of mRNA being produced before release into the cytoplasm where mass protein synthesis occurs. This will suggest that protein synthesis can be regulated in the nucleus as a means for cells to conserve energy. The real life application of such a paradigm is that new efficacious and cost effective methods of translational regulation may target the nucleus since shunting nuclear translation will inhibit further transcription of mRNA and automatically stop the cytoplasmic protein synthesis (the main protein synthesis machinery). This will be of great interest in drug discovery as new therapies will particularly target the nucleus by causing mutations in transcripts to favor NMD. The advancement in nanotechnology will make possible such developments by producing nanoparticles which are site oriented to preferentially target nuclear processes. |
| If nuclear translation actually exists, one of the challenging questions to be addressed is whether non-nuclear proteins can be synthesized in the nucleus? If so, will the synthesized proteins be easily transported out of the nucleus to their target sites since most extra nuclear proteins are usually large in size? Also, could these large non-nuclear proteins which cannot be transported across the nuclear membrane accumulate and have delirious effects to the cell? Can such accumulated non-nuclear proteins be responsible for certain diseases state? All such possible questions may need to be addressed to better understand the implication of such novel paradigm in biology. |
| Conclusion |
| From the era where findings in cell biology was difficult to clearly explain certain cell processes, molecular technologies have now pacify the study of such processes which have laid new paradigms. Though most of these new paradigms have generally being accepted, the concept of nuclear translation still remains unclear to many researchers. Most of these discoveries have provided insights in the understanding of some cellular processes and their mechanisms especially on gene expression and regulation as well as the pathogenesis of certain illnesses. Also, possible targets for therapeutic agents have been identified and are now gaining substantial interest in biomedicine for pharmaceutical and other biomedical applications. Some of these discoveries are being exploited for their usefulness as biosensors for diagnostic tools, selectable markers for gene expression and catalytic material for biotechnological industries. While we hope for fruitful outcomes of such discoveries, we are encouraging high interest in molecular and biotechnological research to explore such theoretical positions for better understanding of certain cellular processes and possible useful applications without mitigating the delirious effects to the society. |
| Glossary |
| Nonsense mediated decay is a scan machinery used by ribosomes to scan messenger RNA (mRNA) for inappropriately placed premature termination codons (PTCs) and destroy faulty messages; Prions are known proteinatious infectious agents capable of causing a disease; Introns are the non-coding sequence of mRNA; Gal-operon is a set of genes involve in the metabolism of galactose; Maxicircle DNA also known as minicircles are small circular DNA molecules that are all linked together by catenation-like rings in a chain forming a giant network of DNA found in the mitochondria of trypanosomes; Cryptogenes are hidden genes whose transcript has been edited extensive with uracyls in the coding region; Guide RNA is a short RNA sequences that base pair with mRNA at 5’ end of the pre-edited region; Retrohoming is a process whereby introns, after their conversion to DNA by reverse transcription, catalyse their insertion into the genome other than the original site; Retrotransposons are mobile genetic elements in humans whose sequence resembles those of retroviruses. |
| Author’s Contribution |
| GA Engwa designed the review, collated the ideas and drafted the manuscript. |
| Acknowledgement |
| My first exposure to these changing paradigms was at my postgraduate studies. I acknowledge the effort of Professor Wilfred Fon Mbacham, my mentor for introducing me to molecular biology research. |
References
|
Tables and Figures at a glance
| Table 1 |
Figures at a glance
 |
 |
 |
 |
| Figure 1 | Figure 2 | Figure 3 | Figure 4 |
 |
 |
 |
 |
| Figure 5 | Figure 6 | Figure 7 | Figure 8 |
Relevant Topics
- Analytical Biochemistry
- Applied Biochemistry
- Carbohydrate Biochemistry
- Cellular Biochemistry
- Clinical_Biochemistry
- Comparative Biochemistry
- Environmental Biochemistry
- Forensic Biochemistry
- Lipid Biochemistry
- Medical_Biochemistry
- Metabolomics
- Nutritional Biochemistry
- Pesticide Biochemistry
- Process Biochemistry
- Protein_Biochemistry
- Single-Cell Biochemistry
- Soil_Biochemistry
Recommended Journals
- Biosensor Journals
- Cellular Biology Journal
- Journal of Biochemistry and Microbial Toxicology
- Journal of Biochemistry and Cell Biology
- Journal of Biological and Medical Sciences
- Journal of Cell Biology & Immunology
- Journal of Cellular and Molecular Pharmacology
- Journal of Chemical Biology & Therapeutics
- Journal of Phytochemicistry And Biochemistry
Article Tools
Article Usage
- Total views: 14142
- [From(publication date):
December-2015 - Jan 15, 2025] - Breakdown by view type
- HTML page views : 9710
- PDF downloads : 4432
